
The Hidden Problem in Dermatology Research
And What We Can Do About It
2026-04-21
Disclosures
Consultant, Advisor, Speaker: Almirall, Bristol Myers Squibb, Castle Biosciences, Checkpoint Therapeutics, EMD Serono, Merck, Pfizer, Sanofi Genzyme
Researcher: Kartos Therapeutics, NeoImmune Tech, Regeneron Pharmaceuticals Inc.
Other (Sterring Committee): Sun Pharmaceuticals Inc.
Individual publicly traded stocks and stock options: Avstera
Who here enjoys discovering new things?
Who here has published in an academic journal?
Who here enjoys data analysis?
Who here enjoys statistics?
Who here has heard about concerns regarding reproducibility in science?
Who here is concerned about public trust in science?
Why This Conversation Matters
Scientific progress depends on credible evidence
Clinical decisions rely on trustworthy results
Research findings influence patient care
Important
How we analyze data influences how evidence is interpreted
One Dominant Framework For Interpreting Evidence
Much of modern biomedical research relies on:
Hypothesis testing
p-values
Statistical significance thresholds
Dichotomous interpretation of results
Important
Conventions for statistical inference influence how evidence is interpreted
Inference
- Using Study Data To Learn What Is Likely True Beyond The Study Itself
An Early Look Into What We Found
Over 4,000 research articles from a prominent dermatology journal
Studies report far more statistical tests than most readers realize
Very little adjustment for multiplicity
Almost no preregistration
Important
The evidence we rely on may be more uncertain than it appears?
Tonight’s Roadmap
Why this conversation matters?
How inferential frameworks shape interpretation of evidence
How the question can be studied empirically
What dermatology literature reveals about analytic structure
Approaches to improve interpretability
Statistical Frameworks And Interpretation
Modern statistical practice combines ideas from multiple historical traditions
These frameworks were developed to answer different scientific questions
Current conventions reflect a blending of historically distinct approaches
Important
Understanding statistical evidence requires understanding the framework being applied
Karl Pearson
Pearson helped establish hypothesis testing as a scientific tool
Developed methods to evaluate whether observed data were consistent with theoretical expectations
1900: introduced the χ² goodness-of-fit test

1857 – 1936
Image Source: Wikipedia
Key idea
Hypothesis testing began as a method for comparing observed vs expected patterns.
R. A. Fisher
Developed the p-value as a measure of strength of evidence against a hypothesis
1925: Statistical Methods for Research Workers
Framed statistical inference as quantifying how surprising the observed data would be under a model

1890 – 1962
Image Source: Wikipedia
Key idea
The p-value was originally proposed as a graded measure of evidence, not a strict decision rule.
Neyman & Pearson
Framed hypothesis testing as a formal decision process between competing hypotheses
1933: Type I / Type II error framework
Introduced fixed decision thresholds and long-run error control

1894 – 1981 (Neyman)
1895 – 1980 (Pearson)
Image sources: https://statistics.berkeley.edu/people/jerzy-neyman. https://mathshistory.st-andrews.ac.uk/Biographies/Pearson_Egon/
Key idea
Neyman and Pearson formalized statistical testing as a decision rule
with prespecified error rates (α and β).
Modern NHST Framework
Statistical evidence is often summarized using p-values
Interpretation often depends on whether the p-value crosses a fixed threshold
Most commonly:
p < 0.05 → “statistically significant”
Important
Modern practice combines:
Fisher
continuous measure of evidence
with
Neyman–Pearson
fixed decision thresholds
A Consequence Of Threshold-Based Inference
When results are judged by whether p < 0.05 —
analytic choices that influence p-values also influence conclusions
Modern studies often involve:
• Multiple outcomes • Multiple models • Multiple analytic decisions
Important
Different analytic paths applied to the same data can produce different conclusions
We Can Measure This
If analytic choices influence conclusions —
then analytic structure should be observable in the published literature
We evaluated statistical reporting patterns across the JAAD corpus to find out.
Studying A Real Clinical Literature
Journal Of The American Academy Of Dermatology
Advantages:
• Diverse study designs
• Frequent use of statistical inference
• Direct relevance to clinical decision-making
Why Statistical Choices Matter for the Evidence Base
Open Science Collaboration
Science 2015
• Replicated 100 top psychology studies
• 97% of originals significant
• 36% significant on replication
• Replication effect sizes were about half as large
Errington et al.
eLife 2021
• Reproducibility Project: Cancer Biology
• 50 replications from 23 high-impact papers
• Effects often smaller on replication
• Only 3% matched or exceeded original effect size
• Identified gaps in methods transparency
Cobey et al.
PLOS Biology 2024
• Survey of 1,630 biomedical researchers
• Researchers from 80+ countries
• 72% perceive a reproducibility crisis
• Publication pressure cited as cause
• Many said novelty favored over verification
Warning
Analytic choices influence which findings enter — and persist in — the literature
Analytic Flexibility and Statistical Significance
When many analytic decisions are possible, statistical conclusions may depend on
which choices are made
Common terminology:
• p-hacking
Selectively exploring analyses
to obtain statistical significance
Analytic Flexibility and Statistical Significance
When many analytic decisions are possible, statistical conclusions may depend on
which choices are made
Common terminology:
• p-hacking
Selectively exploring analyses
to obtain statistical significance
• HARKing
Hypothesizing After Results Are Known
Post-hoc findings framed as pre-planned

Analytic Flexibility and Statistical Significance
When many analytic decisions are possible, statistical conclusions may depend on
which choices are made
Common terminology:
• p-hacking
Selectively exploring analyses
to obtain statistical significance
• HARKing
Hypothesizing After Results Are Known
Post-hoc findings framed as pre-planned
• Researcher Degrees Of Freedom
Multiple defensible analytic choices and each choice shifts the result
Analytic Flexibility and Statistical Significance
When many analytic decisions are possible, statistical conclusions may depend on
which choices are made
Common terminology:

• Researcher Degrees Of Freedom
Multiple defensible analytic choices and each choice shifts the result
Analytic Flexibility and Statistical Significance
When many analytic decisions are possible, statistical conclusions may depend on
which choices are made
Common terminology:

• Researcher Degrees Of Freedom
Multiple defensible analytic choices and each choice shifts the result
• Garden Of Forking Paths
Analytic decisions not prespecified
Data patterns silently guide each fork
Analytic Flexibility and Statistical Significance
When many analytic decisions are possible, statistical conclusions may depend on
which choices are made
Common terminology:
• p-hacking
Selectively exploring analyses
to obtain statistical significance
• HARKing
Hypothesizing After Results Are Known
Post-hoc findings framed as pre-planned
• Researcher Degrees Of Freedom
Multiple defensible analytic choices and each choice shifts the result
• Garden Of Forking Paths
Analytic decisions not prespecified
Data patterns silently guide each fork
Important
Different analytic choices applied to the same data can produce different statistical conclusions.
Multiplicity Increases Probability Of False Positive Findings
When multiple statistical tests are performed, the probability of at least one statistically significant result increases even if no true effect exists
If α = 0.05 for each test:
1 test
Probability of false positive
≈ 5%
10 tests
Probability of ≥1 false positive
≈ 40%
20 tests
Probability of ≥1 false positive
≈ 64%
Important
Even modest numbers of statistical tests substantially increase the probability of at least one statistically significant result.
Multiplicity And Type I Error
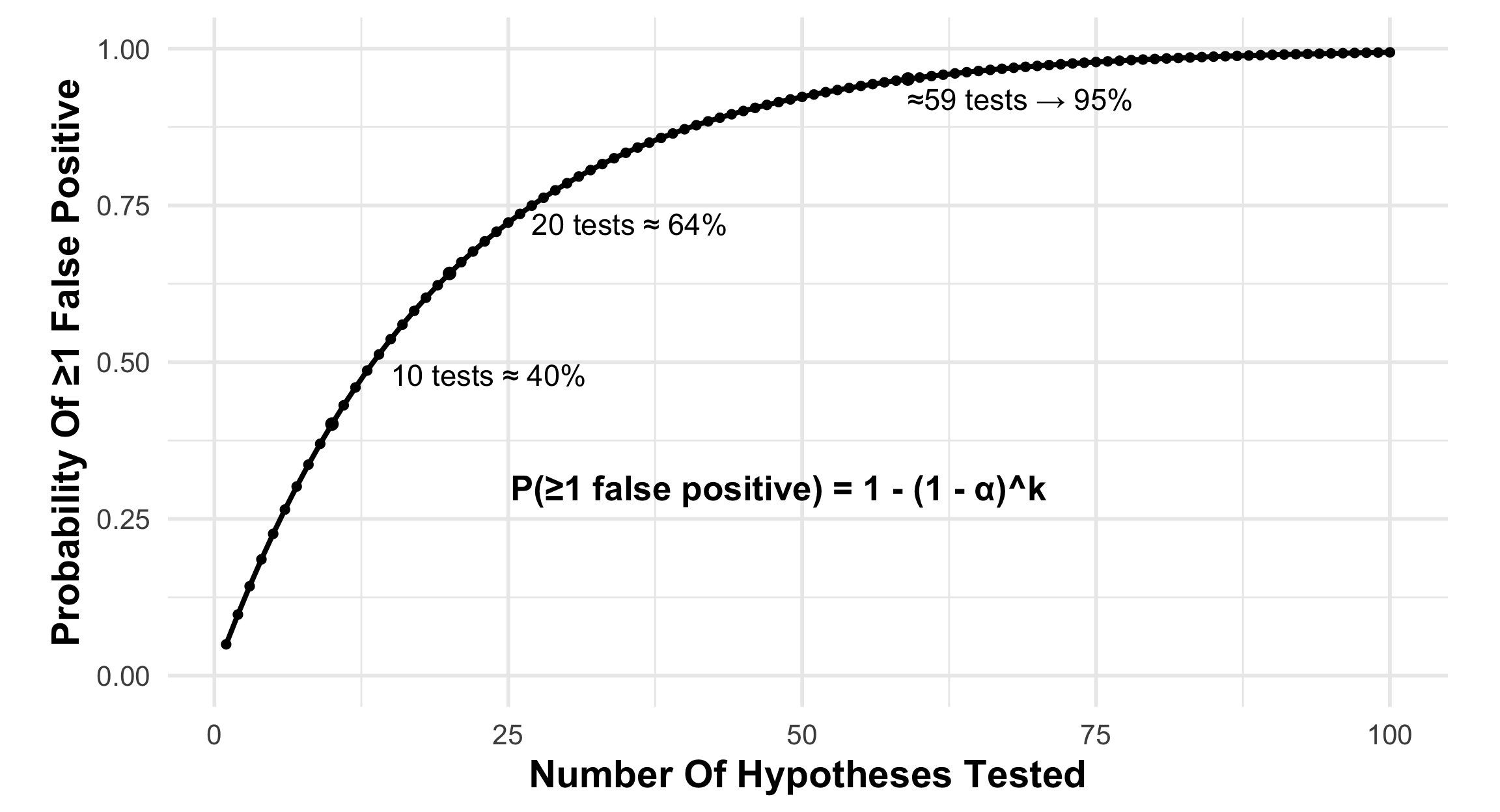
Important
As the number of analytic pathways increases, statistically significant findings become more likely even when no true effect exists.
Where Multiplicity Arises In Biomedical Studies
Multiple outcomes
• Primary endpoints
• Secondary endpoints
• Exploratory endpoints
Multiple models
• Alternative covariate sets
• Different ways of modeling variables (e.g. continuous vs. categorical)
Multiple subgroups
• Age groups
• Disease severity
• Biomarker-defined subgroups
Multiple analytic decisions
• Inclusion criteria
• Missing data handling
• Variable definitions
Important
Multiplicity often arises naturally from reasonable analytic decisions in complex data.
Estimating Analytic Search Space In Dermatology Research
Multiplicity arises because modern studies explore many reasonable analytic pathways
This creates an analytic search space that shapes how statistical evidence should be interpreted
We asked:
How large is the analytic search space in contemporary dermatology research?
We evaluated statistical reporting patterns across the JAAD corpus
From Pilot Review To Scalable Pipeline
Initial manual review (56 articles) suggested a substantial analytic search space
Scaling this evaluation required reproducible methods to extract statistical information from text
We developed a structured pipeline to evaluate analytic search space across a larger corpus
Goal: systematically characterize statistical reporting patterns at scale
Identifying A Reliable Text Source
Statistical information can be extracted from multiple sources

HTML

Identifying A Reliable Text Source
Statistical information can be extracted from multiple sources

HTML
<html class="modern" id="ng-app" lang="en-US" xmlns:ng="http://angularjs.org">
<head>
<style>
@charset "UTF-8";[ng\:cloak],[ng-cloak],[data-ng-cloak],[x-ng-cloak],.ng-cloak,.x-ng-cloak,.ng-hide:not(.ng-hide-animate){display:none !important;}ng\:form{display:block;}.ng-animate-shim{visibility:hidden;}.ng-anchor{position:absolute;}
</style>
<meta content="IE=edge" http-equiv="X-UA-Compatible"/>
<meta charset="utf-8"/>
<meta content="1" name="tdm-reservation"/>
<meta content="https://www-elsevier-com.treadwell.idm.oclc.org/tdm/tdmrep-policy.json" name="tdm-policy"/>
<meta content="width=device-width, initial-scale=1.0, maximum-scale=1.0, user-scalable=no" name="viewport"/>
<title>
Real-world assessment of response to anti-programmed cell death 1 therapy in advanced cutaneous squamous cell carcinoma - ClinicalKey
</title>
<script src="https://js-agent.newrelic.com/nr-spa-1216.min.js">
</script>
<script async="" src="https://cdn.pendo.io/agent/static/b3541d7b-4788-4b73-7811-976020af677d/pendo.js">
</script>
<script type="text/javascript">
;window.NREUM||(NREUM={});NREUM.init={distributed_tracing:{enabled:true},privacy:{cookies_enabled:true},ajax:{deny_list:["bam.nr-data.net"]}};
;NREUM.loader_config={accountID:"1574307",trustKey:"2038175",agentID:"243284150",licenseKey:"94f48af4f8",applicationID:"243284150"}
;NREUM.info={beacon:"bam.nr-data.net",errorBeacon:"bam.nr-data.net",licenseKey:"94f48af4f8",applicationID:"243284150",sa:1}
</path>
<polygon points="29.471 11.883 29.471 15.02 32.609 15.02 32.609 24.435 35.75 24.435 35.75 15.02 38.888 15.02 38.888 11.883 29.471 11.883">
</polygon>
<path d="M15.55139,11.66574c-1.33253,0-2.43215.03784-3.62968,0.118l-0.23665.02446V24.43481H14.8243V19.55419c0,0.01413.47114,0.01413,0.63653,0.01413a4.164,4.164,0,0,0,4.34788-4.00956C19.8087,13.117,18.126,11.66574,15.55139,11.66574ZM14.8243,14.16984l0.65362-.01041c1.06617,0,1.33771.40442,1.33771,1.40753,0,0.71372-.37319,1.54619-2.13379,1.54619H14.8243V14.16984Z">
</path>
<path d="M48,0L0,0.003V35.5574H16.25546L14.7367,40.00025h-5.848v4.44726H39.10989V40.00025H33.25958l-1.518-4.44285H48V0ZM20.25909,40.00025l1.51656-4.44285h4.43907l1.51952,4.44285H20.25909ZM43.5557,31.1109H4.44506V4.44726H43.5557V31.1109Z">
</path>
</symbol>
<symbol id="els-hmds-icon-ppt-2" viewbox="0 0 48 64">
<title>
ppt-2
</title>
<use xlink:href="#icon__ppt-2" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-printer-2" viewbox="0 0 126 128">
<title>
printer-2
</title>
<path d="m97 54h1e1v1e1h-1e1v-1e1zm-6e1 28h52v24h-52v-24zm-1e1 34h72v-44h-72v44zm1e1 -1e2h52v2e1h-52v-2e1zm75 2e1h-13v-3e1h-72v3e1h-13c-7.16 0-13 5.83-13 13v4e1c0 7.17 5.84 13 13 13h5v-1e1h-5c-1.62 0-3-1.37-3-3v-4e1c0-1.63 1.38-3 3-3h98c1.62 0 3 1.37 3 3v4e1c0 1.63-1.38 3-3 3h-5v1e1h5c7.16 0 13-5.83 13-13v-4e1c0-7.17-5.84-13-13-13">
</path>
</symbol>
<symbol id="els-gizmo-icon-publication-set" viewbox="0 0 122 128">
<title>
publication-set
</title>
<path d="m12 57c0-6.08 4.92-11 11-11h17v-2e1h-6c-2.2 0-4 1.8-4 4v6h-7c-3.32 0-6.44 0.78-9.22 2.16 2.44-5.64 7.28-11.86 13.5-17.1 2.34-1.98 5.3-3.06 8.32-3.06h46.4v44.12l8.26-8.26c0.56-0.56 1.14-1.06 1.74-1.54v-44.32h-56.4c-5.38 0-10.62 1.92-14.76 5.4-9.1 7.68-18.84 20.14-18.84 32.1v70.5h41.84l3.12-1e1h-34.96v-49zm97.42 16.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56z">
</path>
</symbol>
<symbol id="els-gizmo-icon-publication-sets" viewbox="0 0 122 128">
<title>
publication-sets
</title>
<path d="m109.42 73.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56zm-104.48 3.46c0-6.08 4.92-11 11-11h17v-2e1h-6c-2.2 0-4 1.8-4 4v6h-7c-3.32 0-6.44 0.78-9.22 2.16 2.44-5.64 7.28-11.86 13.5-17.1 2.34-1.98 5.3-3.06 8.32-3.06h34.4v46.12l1e1 -1e1v-46.12h-44.4c-5.38 0-10.62 1.92-14.76 5.4-9.1 7.68-18.84 20.14-18.84 32.1v60.5h41.84l3.12-1e1h-34.96v-39zm76-10.88l2.26-2.26c2.2-2.2 4.86-3.82 7.74-4.76v-49.1h-44.4c-5.38 0-10.62 1.92-14.76 5.4-1.64 1.38-3.3 2.94-4.92 4.6h54.08v46.12z">
</path>
</symbol>
<symbol id="els-gizmo-icon-radiology" viewbox="0 0 126 128">
<title>
radiology
</title>
<path d="m48 68.5v18.32c0 5.78-2.04 10.74-6.08 14.7-6.48 6.4-15.98 8.44-19.26 8.4-7.18-0.1-10.66-4.46-10.66-13.34 0-20.48 8.76-51.82 20.08-61.68 4.42-3.86 6.6-2.86 7.32-2.52 2.1 0.96 3.94 3.36 5.4 6.44 2.64-3.02 4.4-6.74 4.98-10.74-1.74-2.08-3.8-3.7-6.22-4.8-4-1.82-10.38-2.58-18.04 4.08-15.14 13.2-23.52 49.26-23.52 69.22 0 14.44 7.68 23.16 20.52 23.34h0.24c5.8 0 17.82-3.02 26.2-11.28 5.92-5.84 9.04-13.38 9.04-21.82v-24.84c-2.72 2.12-3.8 2.7-1e1 6.52zm52.5-41.14c-7.66-6.68-14.04-5.9-18.04-4.08-2.44 1.1-4.48 2.74-6.22 4.82 0.58 4 2.34 7.72 4.98 10.74 1.32-2.8 3.92-6.82 6.96-6.82 1.2 0 3.06 0.56 5.76 2.88 11.3 9.84 20.06 41.2 20.06 61.68 0 8.86-3.48 13.24-10.66 13.34-3.34 0-12.78-2-19.26-8.4-4.04-3.96-6.08-8.92-6.08-14.7v-18.32c-6.18-3.8-7.28-4.38-1e1 -6.52v24.84c0 8.44 3.12 15.98 9.06 21.82 8.38 8.26 20.4 11.28 26.2 11.28h0.24c12.82-0.18 20.5-8.9 20.5-23.34 0-19.96-8.38-56.04-23.5-69.22zm-24.1 30.76l14.22 8.76 5.24-8.52-14.24-8.76c-8.4-5.18-13.62-14.54-13.62-24.42v-19.18h-1e1v19.18c0 9.88-5.22 19.24-13.64 24.42l-14.24 8.76 5.24 8.52 14.22-8.76c5.64-3.48 10.22-8.34 13.4-14 3.2 5.66 7.78 10.52 13.42 14z">
</path>
</symbol>
<symbol id="els-gizmo-icon-rainbow" viewbox="0 0 128 128">
<title>
rainbow
</title>
<path d="m105.76 112h-40.8c-5 0-9.08-4.08-9.08-9.1 0-4.7 2.84-8.12 7.78-9.38l4.06-1.02-0.32-4.2c-0.4-5.38 1-9.98 4.06-13.28 2.98-3.24 7.44-5.02 12.5-5.02 8.06 0 14.9 5.8 16.28 13.8l0.66 3.84 3.88 0.3c7.76 0.6 12.96 5.44 12.96 12.06s-5.38 12-11.98 12zm-63.86-44h-26.06c-3.24 0-5.86-2.74-5.86-6.12 0-3.9 3.32-5.9 6.62-6.16l3.88-0.3 0.66-3.84c0.74-4.4 4.36-7.58 8.62-7.58 2.68 0 5 0.94 6.58 2.62 1.66 1.8 2.42 4.4 2.2 7.5l-0.3 4.16 4.04 1.04c1.64 0.44 3.62 1.56 3.62 4.46 0 2.38-1.76 4.22-4 4.22zm67.3 10.48c-3.24-10.14-12.2-17.4-22.84-18.36-6.36-15.62-16.02-18.12-21.5-18.12-6.72 0-12.22 2.92-16.46 8.58-0.46-4.16-2.08-7.88-4.76-10.76-3.48-3.76-8.4-5.82-13.88-5.82-7.92 0-14.82 5.04-17.52 12.38-7.28 1.96-12.24 8.04-12.24 15.5 0 8.88 7.1 16.12 15.84 16.12h26.06c7.7 0 13.98-6.38 13.98-14.22 0-1.8-0.34-3.52-0.9-5.1 3.34-5.88 7-6.68 9.88-6.68 4.16 0 8.02 3.26 11 9.12-4.54 1.3-8.58 3.7-11.72 7.1-4.12 4.44-6.46 10.36-6.74 16.92-7.18 3.2-11.5 9.72-11.5 17.76 0 10.54 8.54 19.1 19.06 19.1h40.8c12.1 0 21.96-9.88 21.96-22 0-10.64-7.6-19.22-18.52-21.52m-44.34-54.48c-7.22 0-13.94 2.32-19.88 6.54 1.62 1.08 3.14 2.36 4.5 3.82 1 1.08 1.86 2.24 2.64 3.46 3.94-2.46 8.22-3.82 12.74-3.82 10.38 0 19.58 7.14 26.04 18.72 5.08 1.04 9.76 3.18 13.78 6.22-7.64-21.2-22.28-34.94-39.82-34.94m-28.24 2.68c8.12-6.92 17.74-10.68 28.24-10.68 25.52 0 45.8 23.16 51.94 56.48 4.38 1.78 8.18 4.42 11.2 7.7-4.54-43.14-30.16-74.18-63.14-74.18-16.52 0-31.18 7.62-42.28 20.82 2.3-0.66 4.7-1 7.18-1 2.36 0 4.66 0.32 6.86 0.86">
</path>
</symbol>
<symbol id="els-gizmo-icon-rainbow-2" viewbox="0 0 128 128">
<title>
rainbow-2
</title>
<path d="m64 66c-15.44 0-28 15.26-28 34h1e1c0-13.24 8.08-24.02 18-24.02s18 10.78 18 24.02h1e1c0-18.74-12.56-34-28-34m0-18c-25.8 0-46 22.84-46 52h1e1c0-23.56 15.82-42 36-42s36 18.44 36 42h1e1c0-29.16-20.2-52-46-52m0-18c-35.88 0-64 30.74-64 7e1h1e1c0-33.64 23.72-6e1 54-6e1s54 26.36 54 6e1h1e1c0-39.26-28.12-7e1 -64-7e1">
</path>
</symbol>
<symbol id="els-gizmo-icon-rar-file" viewbox="0 0 92 128">
<title>
rar-file
</title>
<path d="m34.01 48l3.03-1e1h0.08l2.84 1e1h-5.95zm-0.46-18l-8.55 26h7.19l1.04-4h7.38l0.98 4h7.41l-8.46-26h-6.99m29.6 12h-6.15v-6h5.8c2.44 0 3.17 1.48 3.17 3 0 2.16-1.63 3-2.82 3zm9.21-4.12c0-4.56-3.14-7.88-6.77-7.88h-14.59v26h6v-8h6.21c2.53 0 2.7 2.2 2.88 4.48 0.08 1.26 0.2 1.52 0.52 3.52h6.39c-0.58-2-0.61-3.94-0.7-5.12-0.23-3-1.3-5.2-3.05-5.98 2.12-0.9 3.11-4.54 3.11-7.02m-65.36-1.88h5.8c2.44 0 3.17 1.48 3.17 3 0 2.16-1.63 3-2.82 3h-6.15v-6zm0 12h6.21c2.53 0 2.7 2.2 2.88 4.48 0.08 1.26 0.2 1.52 0.52 3.52h6.39c-0.58-2-0.61-3.94-0.7-5.12-0.23-3-1.3-5.2-3.05-5.98 2.12-0.9 3.11-4.54 3.11-7.02 0-4.56-3.14-7.88-6.77-7.88h-14.59v26h6v-8m-6-38v1e1h8e1v60.96l-26.93 27.04h-43.07v-42h-1e1v52h57.22l32.78-32.92v-75.08h-9e1m42 9e1h1e1v-2e1h2e1v-1e1h-3e1v3e1">
</path>
</symbol>
<symbol id="icon__rationale" viewbox="0 0 47.47561 47.99999">
<title>
rationale
</title>
<rect height="3.94076" width="3.93866" x="25.56554" y="24.30582">
</rect>
<path d="M27.93008,7.75887a5.97685,5.97685,0,0,0-6.3053,6.30409h3.94076c0-1.57732.78822-3.15155,2.36454-3.15155a2.122,2.122,0,0,1,2.32783,2.54327c-0.28854,2.32673-4.69237,3.08043-4.69237,7.19782v1.28988H29.5042V21.15428c0-2.82861,4.729-3.79544,4.729-7.87942C34.23318,10.1222,31.54779,7.75887,27.93008,7.75887Z">
</path>
<path d="M21.7949,48H17.82614V45.2931c-3.58981.39516-8.15716,0.52118-9.96392-1.03365a2.87839,2.87839,0,0,1-1.03365-2.20511V33.05516H0L7.20973,19.144a17.36781,17.36781,0,0,1,1.3868-7.99784C10.98687,5.24361,18.03077-.74294,29.07133.07538A19.60255,19.60255,0,0,1,43.312,8.056c3.85035,5.18543,5.08433,12.00861,3.4757,19.215-0.66969,3.00083-2.88991,4.8862-4.8496,6.55073A17.489,17.489,0,0,0,39.48752,36.169a14.3032,14.3032,0,0,0-2.57767,5.87034v5.265h-3.971l0.01941-5.69581a18.18831,18.18831,0,0,1,3.39808-7.88041,20.791,20.791,0,0,1,3.01274-2.933c1.62793-1.38139,3.16335-2.68748,3.5445-4.38762,1.73564-7.7717-.55767-12.983-2.7887-15.98487A15.85123,15.85123,0,0,0,28.7785,4.03344c-8.66113-.63959-14.57657,3.84495-16.504,8.60292a15.78078,15.78078,0,0,0-1.05724,6.3397l0.183,0.74081-0.35965.68259L6.637,29.0864h4.16039V41.36094c1.3954,0.35315,5.16614.24333,8.71924-.29063l2.27832-.34245V48Z">
</path>
</symbol>
<symbol id="els-hmds-icon-rationale" viewbox="0 0 47.47561 64">
<title>
rationale
</title>
<use xlink:href="#icon__rationale" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-record" viewbox="0 0 96 128">
<title>
record
</title>
<path d="m48 23.62c-10.26 0-19.9 3.99-27.14 11.24s-11.24 16.89-11.24 27.14 4 19.89 11.24 27.14 16.9 11.24 27.14 11.24c10.26 0 19.9-3.99 27.14-11.24 7.26-7.25 11.24-16.89 11.24-27.14s-4-19.89-11.24-27.14-16.88-11.24-27.14-11.24zm0 86.38c-12.82 0-24.88-4.99-33.94-14.06s-14.06-21.12-14.06-33.94 5-24.87 14.06-33.94 21.12-14.06 33.94-14.06 24.88 4.99 33.94 14.06c9.06 9.06 14.06 21.12 14.06 33.94s-5 24.87-14.06 33.94-21.12 14.06-33.94 14.06">
</path>
</symbol>
<symbol id="icon__redo" viewbox="0 0 50 45.3691">
<title>
redo
</title>
<path d="M22.685,0C29.0709,0,34.38174,2.33729,39.403,7.35627,40.857,8.811,43.2475,11.2204,45.37,13.35983v-8.731H50v16.6664H33.33281V16.66562h8.79643c-2.12713-2.14558-4.5376-4.57309-5.9993-6.03479-4.151-4.15024-8.2974-6.002-13.44492-6.002A18.05634,18.05634,0,1,0,40.55509,24.99842h4.684A22.679,22.679,0,1,1,22.685,0Z">
</path>
</symbol>
<symbol id="els-hmds-icon-redo" viewbox="0 0 50 64">
<title>
redo
</title>
<use xlink:href="#icon__redo" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="icon__reduce" viewbox="0 0 48 48.00779">
<title>
reduce
</title>
<polygon points="48 3.356 44.64 0 29.696 14.942 29.696 1.215 24.95 1.215 24.95 23.058 46.795 23.058 46.795 18.311 33.045 18.311 48 3.356">
</polygon>
<polygon points="1.203 29.708 14.942 29.708 0 44.65 3.358 48.008 18.3 33.066 18.3 46.805 23.05 46.805 23.05 24.958 1.203 24.958 1.203 29.708">
</polygon>
</symbol>
<symbol id="els-hmds-icon-reduce" viewbox="0 0 48 64">
<title>
reduce
</title>
<use xlink:href="#icon__reduce" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-refresh" viewbox="0 0 112 128">
<title>
refresh
</title>
<path d="m74 6e1h36v-36h-1e1v18.86c-4.58-4.62-9.75-9.83-12.89-12.97-10.84-10.84-22.32-15.89-36.11-15.89-27.02 0-49 21.98-49 49s21.98 49 49 49c25.33 0 46.2-19.32 48.72-44h-10.09c-2.46 19.14-18.82 34-38.63 34-21.5 0-39-17.5-39-39s17.5-39 39-39c11.12 0 20.07 4 29.04 12.96 3.16 3.16 8.36 8.41 12.96 13.04h-19v1e1">
</path>
</symbol>
<symbol id="els-gizmo-icon-remove-document" viewbox="0 0 92 128">
<title>
remove-document
</title>
<path d="m29 4e1h34v1e1h-34v-1e1zm14 6e1h1e1v-2e1h2e1v-1e1h-3e1v3e1m38-19.04l-26.93 27.04h-43.07v-88h7e1v60.96zm-8e1 -70.96v108h57.22l32.78-32.92v-75.08h-9e1">
</path>
</symbol>
<symbol id="els-gizmo-icon-repeat" viewbox="0 0 111 128">
<title>
repeat
</title>
<path d="m102.24 42.91-7.16 7.16c2.12 3.29 3.38 6.96 3.38 11.16 0 11.58-9.42 20.77-21 20.77h-44.2l13.38-13.13-7.08-6.96-25.44 25.51 25.44 25.49 7.08-7.29-13.4-13.62h44.22c17.1 0 31-13.67 31-30.77 0-6.96-2.34-13.14-6.22-18.32m-89.78 18.32c0-11.58 9.42-21.23 21-21.23h44.22l-13.38 13.61 7.08 7.18 25.44-25.39-25.44-25.43-7.08 6.85 13.4 13.18h-44.24c-17.1 0-31 14.14-31 31.23 0 6.97 2.34 13.51 6.24 18.69l7.16-7.22c-2.14-3.29-3.4-7.26-3.4-11.47">
</path>
</symbol>
<symbol id="els-gizmo-icon-replay" viewbox="0 0 108 128">
<title>
replay
</title>
<path d="m59 16c-13.79 0-25.27 5.05-36.11 15.89-3.14 3.14-8.31 8.35-12.89 12.97v-18.86h-1e1v36h36v-1e1h-19c4.6-4.64 9.8-9.88 12.96-13.04 8.96-8.96 17.92-12.96 29.04-12.96 21.5 0 39 17.5 39 39s-17.5 39-39 39c-19.8 0-36.13-14.86-38.6-34h-10.12c2.52 24.68 23.39 44 48.72 44 27.02 0 49-21.98 49-49s-21.98-49-49-49">
</path>
</symbol>
<symbol id="els-gizmo-icon-research-area" viewbox="0 0 104 128">
<title>
research-area
</title>
<path d="m66 78h26v26h-26v-26zm-1e1 36h46v-46h-46v46zm-44-36h26v26h-26v-26zm-1e1 36h46v-46h-46v46zm54-1e2h46v46h-46zm-44 1e1h26v26h-26v-26zm-1e1 36h46v-46h-46v46z">
</path>
</symbol>
<symbol id="els-gizmo-icon-research-area-edit" viewbox="0 0 126 128">
<title>
research-area-edit
</title>
<path d="m113.42 73.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56zm-54.48 18.58v-16.12h16.12l1e1 -1e1h-36.12v36.12zm-54-16.12h26v26h-26v-26zm-1e1 36h46v-46h-46v46zm54-1e2h46v46h-46zm-44 1e1h26v26h-26v-26zm-1e1 36h46v-46h-46v46z">
</path>
</symbol>
<symbol id="els-gizmo-icon-research-areas" viewbox="0 0 104 128">
<title>
research-areas
</title>
<path d="m66 78h26v26h-26v-26zm-1e1 36h46v-46h-46v46zm-54-46h46v46h-46zm54-54h46v46h-46zm-44 1e1h26v26h-26v-26zm-1e1 36h46v-46h-46v46z">
</path>
</symbol>
<symbol id="els-gizmo-icon-research-areas-edit" viewbox="0 0 126 128">
<title>
research-areas-edit
</title>
<path d="m113.42 73.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56zm-28.36-7.54h-36.12v36.12l1e1 -1e1v-16.12h16.12zm-90.12 0h46v46h-46zm54-54h46v46h-46zm-44 1e1h26v26h-26v-26zm-1e1 36h46v-46h-46v46z">
</path>
</symbol>
<symbol id="els-gizmo-icon-researcher" viewbox="0 0 120 128">
<title>
researcher
</title>
<path d="m107.42 73.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56zm-60.16 24.26l9.38-9.38c-3.06-0.26-6.28-0.42-9.66-0.42-30.88 0-48.88 11.22-51.04 31.74l-0.92 10.26h10.04l0.84-9.28c1.98-18.78 23.34-22.92 41.1-22.92 0.08-0.02 0.18 0 0.26 0zm-0.28-70.08c9.72 0 18.24 8.68 18.24 18.58 0 13.68-7.84 23.98-18.24 23.98s-18.24-10.32-18.24-23.98c0-9.9 8.52-18.58 18.24-18.58zm0 52.28c15.96 0 28-14.48 28-33.66 0-15.36-12.82-28.34-28-28.34s-28 12.98-28 28.34c0 19.18 12.04 33.66 28 33.66z">
</path>
</symbol>
<symbol id="els-gizmo-icon-researcher-profile-needs-action" viewbox="0 0 128 128">
<title>
researcher-profile-needs-action
</title>
<path d="m51.96 9.72c9.72 0 18.24 8.68 18.24 18.58 0 13.68-7.84 23.98-18.24 23.98s-18.24-10.32-18.24-23.98c0-9.9 8.52-18.58 18.24-18.58zm0 52.28c15.96 0 28-14.48 28-33.66 0-15.36-12.82-28.34-28-28.34s-28 12.98-28 28.34c0 19.18 12.04 33.66 28 33.66zm36.04 4e1h1e1v1e1h-1e1zm1e1 -28h-1e1v6l2 18h6l2-18zm-5 44.2c-13.92 0-25.2-11.28-25.2-25.2s11.28-25.2 25.2-25.2 25.2 11.28 25.2 25.2-11.28 25.2-25.2 25.2zm0-60.2c-19.32 0-35 15.68-35 35s15.68 35 35 35 35-15.68 35-35-15.68-35-35-35zm-92.08 43.74l-0.92 10.26h10.04l0.84-9.28c1.92-18.28 22.22-22.68 39.64-22.92 0.98-3.44 2.38-6.7 4.12-9.74-6.64-0.14-12.94 0.12-19.5 1.24-20.58 3.52-32.5 14-34.22 30.44z">
</path>
</symbol>
<symbol id="els-gizmo-icon-researcher-profile-updated" viewbox="0 0 128 128">
<title>
researcher-profile-updated
</title>
<path d="m95 118c-6.68 0-12.6-3.14-16.44-8h9.44v-1e1h-26v26h1e1v-8.26c5.68 6.3 13.88 10.26 23 10.26 15.38 0 28.16-11.28 30.56-26h-10.18c-2.26 9.16-10.52 16-20.38 16zm23-5e1v8.26c-5.68-6.3-13.88-10.26-23-10.26-15.38 0-28.16 11.28-30.56 26h10.18c2.26-9.16 10.52-16 20.38-16 6.68 0 12.6 3.14 16.44 8h-9.44v1e1h26v-26h-1e1zm-66.04 2c-30.88 0-48.88 11.22-51.04 31.74l-0.92 10.26h10.04l0.84-9.28c1.98-18.78 23.34-22.92 41.1-22.92 2.52 0 5.12 0.08 7.72 0.28 1.62-3.36 3.68-6.44 6.12-9.18-4.3-0.58-8.9-0.9-13.86-0.9zm0-60.28c9.72 0 18.24 8.68 18.24 18.58 0 13.68-7.84 23.98-18.24 23.98s-18.24-10.32-18.24-23.98c0-9.9 8.52-18.58 18.24-18.58zm0 52.28c15.96 0 28-14.48 28-33.66 0-15.36-12.82-28.34-28-28.34s-28 12.98-28 28.34c0 19.18 12.04 33.66 28 33.66z">
</path>
</symbol>
<symbol id="els-gizmo-icon-retweet" viewbox="0 0 123 128">
<title>
retweet
</title>
<path d="m113.64 70.3-13.64 13.41v-42.24c0-11.6-8.94-21.47-20.54-21.47h-36c-0.88 0-1.76 0.07-2.6 0.18l9.82 9.82h28.78c6.08 0 10.54 5.39 10.54 11.47v42.19l-13.14-13.36-6.94 7.07 25.5 25.46 25.38-25.46-7.16-7.07m-70.18 21.7c-6.08 0-11.46-4.46-11.46-10.53v-42.21l13.6 13.38 7.2-7.07-25.4-25.46-25.42 25.45 6.84 7.08 13.18-13.39v42.22c0 11.59 9.86 20.53 21.46 20.53h36c0.88 0 1.76-0.07 2.62-0.18l-9.82-9.82h-28.8">
</path>
</symbol>
<symbol id="els-gizmo-icon-rewind" viewbox="0 0 86 128">
<title>
rewind
</title>
<path d="m40.54 101.07l-39.54-39.53 39.54-39.54 7.06 7.07-32.46 32.47 32.46 32.46-7.06 7.07m38 0l-39.54-39.53 39.54-39.54 7.06 7.07-32.46 32.47 32.46 32.46-7.06 7.07">
</path>
</symbol>
<symbol id="els-gizmo-icon-right" viewbox="0 0 104 128">
<title>
right
</title>
<path d="m43.96 19.74l40.26 40.26h-84.22v1e1h84.22l-40.26 40.26 7.08 7.06 52.32-52.32-52.32-52.32z">
</path>
</symbol>
<symbol id="icon__rotate" viewbox="0 0 43.34448 56.34127">
<title>
rotate
</title>
<path d="M17.433,38.18059l-1.41833,1.41833-1.41833,1.41833,2.27487,2.275,2.275,2.275-0.0085.05237A17.47677,17.47677,0,0,1,4.014,28.151V28.14973A17.7,17.7,0,0,1,14.9107,11.8281l-1.536-3.71074A21.63421,21.63421,0,0,0,0,28.14863V28.151A21.47416,21.47416,0,0,0,5.71966,42.64423a21.96045,21.96045,0,0,0,12.78883,6.91892l-0.00521.03276-1.95353,1.95353L14.59635,53.503l1.41915,1.41915,1.41915,1.41915,4.53973-4.53973,4.53986-4.53986-4.54069-4.54041Z">
</path>
<path d="M37.62482,13.697A21.95985,21.95985,0,0,0,24.836,6.77826l0.00521-.0329,1.95353-1.95353,1.95353-1.95353L27.32911,1.41915,25.91,0,21.3701,4.53986,16.83037,9.07959l4.54055,4.54055,4.54055,4.54055,1.41847-1.41833,1.41833-1.41847-2.275-2.27487-2.275-2.27487,0.0085-.05237A17.4765,17.4765,0,0,1,39.33049,28.19031v0.00123A17.7,17.7,0,0,1,28.43378,44.51331l1.536,3.71074A21.6345,21.6345,0,0,0,43.34448,28.19264V28.19031A21.47389,21.47389,0,0,0,37.62482,13.697Z">
</path>
<rect height="8.32302" transform="translate(-13.81906 24.05156) rotate(-45)" width="8.32302" x="17.96176" y="24.54535">
</rect>
</symbol>
<symbol id="els-hmds-icon-rotate" viewbox="0 0 43.34448 64">
<title>
rotate
</title>
<use xlink:href="#icon__rotate" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-rows" viewbox="0 0 110 128">
<title>
rows
</title>
<path d="m12 82h86v18h-86v-18zm0-28h86v18h-86v-18zm0-28h86v18h-86v-18zm-1e1 84h106v-94h-106v94z">
</path>
</symbol>
<symbol id="icon__ruler" viewbox="0 0 22.9385 49.5977">
<title>
ruler
</title>
<path d="M0,0V49.5977H22.9385V0H0ZM3.918,45.1973V4.3994H19.0205V8.6543H10.5772v3.9336h8.4433v3.1455H10.5772v3.9307h8.4433v3.1455H8.1182v3.9307H19.0205v3.1464H10.5772v3.9317h8.4433v3.1455H10.5772v3.9307h8.4433v4.3027H3.918Z">
</path>
</symbol>
<symbol id="els-hmds-icon-ruler" viewbox="0 0 22.9385 64">
<title>
ruler
</title>
<use xlink:href="#icon__ruler" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="icon__ruler-rotate" viewbox="0 0 42 41.4023">
<title>
ruler-rotate
</title>
<path d="M6.31285,7.52618A1.99857,1.99857,0,0,1,8.22328,5.44723h7.65755L13.45291,7.91581l1.28284,1.30365,4.61793-4.607L14.73575,0,13.45291,1.24325,15.88179,3.633H8.22328A3.81085,3.81085,0,0,0,4.49862,7.52618v6.53028a3.96665,3.96665,0,0,0,.03274.4749l1.7815-1.78155V7.52618Z">
</path>
<path d="M2.02954,12.75171l2.474,2.43193V7.52117A3.81209,3.81209,0,0,1,8.22872,3.627H14.759a3.91941,3.91941,0,0,1,.47337.03269l-1.781,1.78155H8.22872a2,2,0,0,0-1.91139,2.08v7.6545l2.38229-2.424L9.96119,14.0346,5.33286,18.65151,0.73133,14.0346Z">
</path>
<path d="M22,0.4023v22H0v19H42v-41H22Zm17,8H31v3h8v2H31v2h8v3H29v4H39v3H31v3h8v1H31v5h8v3H26v-34H39v5Z">
</path>
</symbol>
<symbol id="els-hmds-icon-ruler-rotate" viewbox="0 0 42 64">
<title>
ruler-rotate
</title>
<use xlink:href="#icon__ruler-rotate" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-save-file" viewbox="0 0 104 128">
<title>
save-file
</title>
<path d="m74 54h-8v-22h-1e1v22h-26v-3e1h44zm-54-4e1v5e1h64v-39.45l1e1 9.96v71.49h-84v-92h-1e1v102h104v-85.64l-16-16.36z">
</path>
</symbol>
<s
</title>
<path d="m19.22 76.91c-5.84-5.84-9.05-13.6-9.05-21.85s3.21-16.01 9.05-21.85c5.84-5.83 13.59-9.05 21.85-9.05 8.25 0 16.01 3.22 21.84 9.05 5.84 5.84 9.05 13.6 9.05 21.85s-3.21 16.01-9.05 21.85c-5.83 5.83-13.59 9.05-21.84 9.05-8.26 0-16.01-3.22-21.85-9.05zm80.33 29.6l-26.32-26.32c5.61-7.15 8.68-15.9 8.68-25.13 0-10.91-4.25-21.17-11.96-28.88-7.72-7.71-17.97-11.96-28.88-11.96s-21.17 4.25-28.88 11.96c-7.72 7.71-11.97 17.97-11.97 28.88s4.25 21.17 11.97 28.88c7.71 7.71 17.97 11.96 28.88 11.96 9.23 0 17.98-3.07 25.13-8.68l26.32 26.32 7.03-7.03">
</path>
</symbol>
<symbol id="els-gizmo-icon-search-document" viewbox="0 0 110 128">
<title>
search-document
</title>
<path d="m69 108c-10.5 0-19-8.5-19-19s8.5-19 19-19 19 8.5 19 19-8.5 19-19 19zm23.72-2.34c3.32-4.72 5.28-10.46 5.28-16.66 0-16.02-12.98-29-29-29s-29 12.98-29 29 12.98 29 29 29c6.2 0 11.94-1.96 16.66-5.28l14.82 14.82 7.08-7.08-14.84-14.8zm-80.72-3.66v-49c0-6.08 4.92-11 11-11h17v-2e1h-6c-2.2 0-4 1.78-4 4v6h-7c-3.32 0-6.44 0.78-9.22 2.16 2.46-5.62 7.28-11.86 13.5-17.1 2.34-1.98 5.3-3.06 8.32-3.06h46.4v40.18c3.64 1.36 7 3.26 1e1 5.62v-55.8h-56.4c-5.38 0-10.62 1.92-14.76 5.4-9.1 7.68-18.84 20.14-18.84 32.1v70.5h37.8c-2.36-3-4.26-6.36-5.62-1e1h-22.18z">
</path>
</symbol>
<symbol id="els-gizmo-icon-secondary-result" viewbox="0 0 117 128">
<title>
secondary-result
</title>
<path d="m2e1 1e1h68v1e1h-68zm0 22h68v1e1h-68zm-18-22h1e1v1e1h-1e1zm0 22h1e1v1e1h-1e1zm0 22h1e1v1e1h-1e1zm0 22h1e1v1e1h-1e1zm18-22v1e1h22.98c1.64-3.7 3.86-7.06 6.54-1e1h-29.52zm19.96 22h-19.96v1e1h20.48c-0.44-2.26-0.68-4.6-0.68-7 0-1.02 0.08-2 0.16-3zm18.04 3c0-10.5 8.5-19 19-19s19 8.5 19 19-8.5 19-19 19-19-8.5-19-19zm57.54 31.46l-14.82-14.82c3.32-4.7 5.28-10.44 5.28-16.64 0-16.02-12.98-29-29-29s-29 12.98-29 29 12.98 29 29 29c6.2 0 11.94-1.96 16.66-5.28l14.82 14.82 7.06-7.08z">
</path>
</symbol>
<symbol id="els-gizmo-icon-selection-panel-add" viewbox="0 0 128 128">
<title>
selection-panel-add
</title>
<path d="m24 5e1h1e1v1e1h-1e1zm18 0h36v1e1h-36zm-18-2e1h1e1v1e1h-1e1zm18 0h36v1e1h-36zm48 72v1e1h1e1v-1e1h1e1v-1e1h-1e1v-1e1h-1e1v1e1h-1e1v1e1zm5 16.2c-11.7 0-21.2-9.5-21.2-21.2s9.5-21.2 21.2-21.2 21.2 9.5 21.2 21.2-9.5 21.2-21.2 21.2zm0-52.2c-17.12 0-31 13.88-31 31s13.88 31 31 31 31-13.88 31-31-13.88-31-31-31zm-53 4v1e1h17.72c1.78-3.7 4.12-7.06 6.9-1e1h-24.62zm-3e1 22v-74h78v40.16c1.64-0.2 3.3-0.36 5-0.36s3.36 0.14 5 0.36v-50.16h-98v94h54.16c-0.22-1.64-0.36-3.3-0.36-5s0.14-3.36 0.36-5h-44.16zm12-22h1e1v1e1h-1e1z">
</path>
</symbol>
<symbol id="els-gizmo-icon-selection-panel-remove" viewbox="0 0 128 128">
<title>
selection-panel-remove
</title>
<path d="m24 7e1h1e1v1e1h-1e1zm18-4e1h36v1e1h-36zm-18 0h1e1v1e1h-1e1zm18 2e1h36v1e1h-36zm-18 0h1e1v1e1h-1e1zm-12 42v-74h78v40.16c1.64-0.2 3.3-0.36 5-0.36s3.36 0.14 5 0.36v-50.16h-98v94h54.16c-0.22-1.64-0.36-3.3-0.36-5s0.14-3.36 0.36-5h-44.16zm3e1 -22v1e1h17.72c1.78-3.7 4.12-7.06 6.9-1e1h-24.62zm53-4c-17.12 0-31 13.88-31 31s13.88 31 31 31 31-13.88 31-31-13.88-31-31-31zm0 52.2c-11.7 0-21.2-9.5-21.2-21.2s9.5-21.2 21.2-21.2 21.2 9.5 21.2 21.2-9.5 21.2-21.2 21.2zm-13.44-14.04l6.28 6.28 7.16-7.16 7.16 7.16 6.28-6.28-7.16-7.16 7.16-7.16-6.28-6.28-7.16 7.16-7.16-7.16-6.28 6.28 7.16 7.16z">
</path>
</symbol>
<symbol id="els-gizmo-icon-send" viewbox="0 0 125 128">
<title>
send
</title>
<path d="m113.35 14.88l-111.54 22.73 25.11 22.76 9.03-5.32-12.79-11.58 73.84-15.05-64 37.68v50.57l23.01-18.24-7.57-6.76-5.44 4.31v-19.9l36.34 32.4 43.56-86.17c-2.33-1.81-3.09-2.41-9.55-7.43zm-8.18 20.33l-28.9 57.15-27.12-24.17 56.02-32.98z">
</path>
</symbol>
<symbol id="els-gizmo-icon-settings" viewbox="0 0 120 128">
<title>
settings
</title>
<path d="m60.11 42c-11.58 0-21 9.42-21 21s9.42 21 21 21 21-9.42 21-21-9.42-21-21-21zm0 1e1c6.06 0 11 4.94 11 11s-4.94 11-11 11-11-4.94-11-11c0-2.94 1.14-5.7 3.22-7.78s4.85-3.22 7.78-3.22zm-11.95-46-5.06 12.7-7.49 3.57-13.16-4.01-14.7 18.31 6.87 11.87l-1.84 8-11.37 7.73 5.25 22.86 13.62 2.06 5.16 6.44-1 13.65 21.21 10.12 10.11-9.3h8.32l10.11 9.3 21.21-10.13-1-13.65 5.16-6.43 13.62-2.06 5.25-22.86-11.38-7.74-1.84-8 6.87-11.87-14.7-18.31-13.16 4-7.49-3.56-5.05-12.69h-23.52zm6.79 1e1h9.96l3.99 10.03 14.53 6.94 10.41-3.17 6.18 7.69-5.41 9.35 3.6 15.66 8.96 6.1-2.2 9.57-10.75 1.63-10.07 12.54 0.79 10.77-8.96 4.26-8-7.37h-16.11l-8 7.36-8.96-4.27 0.79-10.76-10.07-12.54-10.75-1.63-2.2-9.57 8.96-6.1 3.6-15.66-5.41-9.35 6.18-7.7 10.41 3.17 14.53-6.92 3.99-10.03z"
<span data-once-text="search_in">
in this
</span>
<button class="j-metrics-click" data-event-label="QuickInContentSearch" data-event-value="InContent" data-metadata-srctype="journal" expand-attributes="data-metadata-searchTerm|data-metadata-srctype\|journal" ng-click="contentSearch()">
<!-- ngIf: srctype === 'book' -->
<!-- ngIf: srctype === 'journal' -->
<span class="ng-scope" data-once-text="Messages.shared_content_journal_article_short" ng-if="srctype === 'journal'">
Article
</span>
<!-- end ngIf: srctype === 'journal' -->
<!-- ngIf: srctype === 'emc' -->
<!-- ngIf: srctype !== 'book' && srctype !== 'journal' && srctype !== 'emc' -->
</button>
<!-- ngIf: srctype === 'journal' || srctype === 'emc' -->
<span class="ng-scope" ng-if="srctype === 'journal' || srctype === 'emc'">
,
</span>
<!-- end ngIf: srctype === 'journal' || srctype === 'emc' -->
<!-- ngIf: srctype === 'book' -->
<!-- ngIf: srctype === 'journal' -->
<span class="ng-scope" ng-if="srctype === 'journal'">
<button class="j-metrics-click" data-event-label="QuickInContentSearch" data-event-value="ParentSource" data-metadata-srctype="journal" data-once-text="Messages.content_journal_issue" expand-attributes="data-metadata-searchTerm|data-metadata-srctype\|journal" ng-click="parentSearch()">
Issue
</button>
,
<span data-once-text="Messages.content_search_or">
or
</span>
<button class="j-metrics-click" data-event-label="QuickInContentSearch" data-event-value="AllJournals" data-metadata-srctype="journal" data-once-text="Messages.shared_content_journal" expand-attributes="data-metadata-searchTerm|data-metadata-srctype\|journal" ng-click="journalSearch()">
Journal
</button>
</span>
<!-- end ngIf: srctype === 'journal' -->
<!-- ngIf: srctype === 'emc' -->
</p>
<!-- end ngIf: formFactor > FORM_FACTORS.MOBILE_LANDSCAPE -->
<div class="ref-text">
<p class="ng-binding" ng-bind-html="refInfo.citationText" ng-hide="refLoading">
</p>
<p class="ng-hide" data-once-text="Messages.reference_loading" ng-show="refLoading">
Loading reference...
</p>
<ul ng-hide="refLoading">
<li ng-show="viewInRefsFn">
<button class="c-link c-link--pane" data-once-text="Messages.reference_view" ng-click="viewInRefsFn({scrollTo: '#' + refInfo.id});close()">
View in References
</button>
</li>
<li class="ng-hide" ng-show="refInfo.doi">
<a class="c-link c-link--pane" data-once-text="Messages.reference_cross" target="_blank">
Cross Reference
</a>
</li>
<li class="ng-hide" data-once-text="Messages.reference_related" ng-show="refInfo.relatedArticles">
Related Articles
</li>
</ul>
</div>
<p class="close">
<button ck-tooltip="Messages.reference_close" class="j-reference-close icon icon-cross-white ng-scope" ng-click="close()">
<span class="visuallyhidden" data-once-text="Messages.reference_close">
Close
</span>
</button>
</p>
</div>
</div>
</div>
<div ck-outline="" class="x-outline-menu j-outline-menu outline-menu ng-isolate-scope" content-type="pgs" hide-eid="true" id-key="sectionid" name-key="subtitle" ng-class="{open: open}" ng-show="XocsCtrl.outlineData.length > 0" outline-data="XocsCtrl.outlineData" stop-propagation="click" update-fn="scrollToFunc">
<h3 class="visuallyhidden" data-once-text="Messages.outline_menu_go_to" id="outline_menu_go_to">
Go to:
</h3>
<div class="outline-container">
<button aria-expanded="false" aria-labelledby="outline_menu_go_to" class="j-outline-header trigger" ng-click="toggleOutline(false)">
<!-- ngIf: contentType === 'BK' -->
<!-- ngIf: contentType !== 'BK' -->
<span class="ng-scope" data-once-text="Messages.outline_menu_outline" ng-if="contentType !== 'BK'">
Outline
</span>
<!-- end ngIf: contentType !== 'BK' -->
<span class="icon icon-arrow-down">
</span>
</button>
<ol aria-hidden="true" class="j-outline-pane pane">
<!-- ngRepeat: item in outlineData track by $index -->
<li class="ng-scope" ng-class="{'active': item.subActive}" ng-repeat="item in outlineData track by $index">
<!-- ngIf: item.eid && !hideEid -->
<!-- ngIf: item[idKey] -->
<a ck-scroll-to="" class="ng-scope" href="hl0000465" ng-click="toggleOutline(item.childrenStore, $index)" ng-href="hl0000465" ng-if="item[idKey]" tabindex="-1" update-fn="select">
<span class="" ng-bind-html="item[nameKey]">
References
</span>
</a>
<!-- end ngIf: item[idKey] -->
<!-- ngIf: item.childrenStore -->
</li>
<!-- end ngRepeat: item in outlineData track by $index -->
</ol>
</div>
</div>
<nav>
<!-- ngIf: XocsCtrl.outlineData -->
<div ck-outline-highlight="outline-highlight" class="outline-container ng-scope" current-section="false" ng-class="{disabledOutline: ContentCtrl.showPaywall}" ng-if="XocsCtrl.outlineData" outline-content='[{"sectionid":"hl0000465","chapternum":"1","subtitle":"References","level":0}]'>
<div class="outline-container__arrow-container">
<div class="outline-container__arrow up">
<span class="icon icon-arrow-up-blue">
</span>
</div>
</div>
<div ck-content-outline="" class="outline-screen ng-isolate-scope" content-type="ContentCtrl.srctype" eid="ContentCtrl.eid" outline-content="outlineContent" scroll-fn="scrollToFunc">
<ul ng-class="{'o-plain-list': contentType === 'core_planning_guide', 'c-content-tabbed__sub-nav-list': contentType === 'core_planning_guide'}">
<!-- ngRepeat: item in outlineContent track by $index -->
<li class="ng-scope" ng-repeat="item in outlineContent track by $index">
<div class="level1" ng-class="'level' + (item.level + 1)">
<!-- ngIf: !item.externalLink -->
<a ck-scroll-to="" class="c-link--nav ng-binding ng-scope" href="#!/content/journal/1-s2.0-S0190962221001973?scrollTo=%23hl0000465" ng-bind-html="item.subtitle || item.itemtitle || item.text || item.outlineLabel" ng-click="fixedHeaderData.currentSection = $event.target.attributes.href.value" ng-if="!item.externalLink" scroll-to-id="hl0000465" update-fn="scrollFn">
References
</a>
<!-- end ngIf: !item.externalLink -->
<!-- ngIf: item.externalLink -->
</div>
</li>
<!-- end ngRepeat: item in outlineContent track by $index -->
</ul>
</div>
<div class="outline-container__arrow-box">
<div class="outline-container__arrow down">
<span class="icon icon-arrow-down-blue">
</span>
</div>
</div>
</div>
<!-- end ngIf: XocsCtrl.outlineData -->
</nav>
<article class="xocs-content__article">
<div class="xocs-content__article-container">
<header class="article-header">
<!-- ngInclude: 'modules/content/partials/' + ContentCtrl.srctype + '-header-partial.html' -->
<div class="ng-scope" ng-include="'modules/content/partials/' + ContentCtrl.srctype + '-header-partial.html'">
<div class="ng-scope">
<p class="content-type ng-binding">
Full Text Article
</p>
<h1>
<span class="ng-binding" ng-bind-html="XocsCtrl.articleTitle">
Real-world assessment of response to anti-programmed cell death 1 therapy in advanced cutaneous squamous cell carcinoma
</span>
<!-- ngIf: XocsCtrl.rssLink -->
<a ck-tooltip="Messages.toolbar_rss" class="x-rss rss icon icon-rss-blue-large ng-scope" href="https://cdn.clinicalkey.com/rss/issue/01909622.xml" ng-href="https://cdn.clinicalkey.com/rss/issue/01909622.xml" ng-if="XocsCtrl.rssLink" target="_blank">
<span class="visuallyhidden">
RSS
</span>
</a>
<!-- end ngIf: XocsCtrl.rssLink -->
<!-- ngIf: context.toolbarData.useOptions.showPDF -->
<a action="download" ck-analytics-click="XocsCtrl.pdfAnalytics" ck-pdf-download="" ck-tooltip="Messages.toolbar_pdf" class="x-pdf j-pdf-trigger icon icon-pdf-red-large pdf ng-scope" eid="1-s2.0-S0190962221001973\01909622/S0190962221X00096/S0190962221001973/main.pdf" href="/service/content/pdf/watermarked/1-s2.0-S0190962221001973.pdf?locale=en_US&searchIndex=" index-override="" ng-if="context.toolbarData.useOptions.showPDF" target="_blank">
<span class="visuallyhidden">
Download PDF
</span>
</a>
<!-- end ngIf: context.toolbarData.useOptions.showPDF -->
</h1>
<!-- ngIf: XocsCtrl.aipStatus === 'S5' || XocsCtrl.aipStatus === 'S100' || XocsCtrl.aipStatus === 'S200' -->
<!-- ngIf: XocsCtrl.embargo -->
<ul class="author-source-list ng-binding" ng-bind-html="XocsCtrl.authorsHtml">
<li>
<a href="#!/search/Shalhout%20Sophia Z./%7B%22type%22:%22author%22%7D">
Sophia Z. Shalhout
<span>
PhD
</span>
</a>
</li>
<li>
,
<a href="#!/search/Park%20Jong Chul/%7B%22type%22:%22author%22%7D">
Jong Chul Park
<span>
MD
</span>
</a>
</li>
<li>
,
<a href="#!/search/Emerick%20Kevin S./%7B%22type%22:%22author%22%7D">
Kevin S. Emerick
<span>
MD
</span>
</a>
</li>
<li>
,
<a href="#!/search/Sullivan%20Ryan J./%7B%22type%22:%22author%22%7D">
Ryan J. Sullivan
<span>
MD
</span>
</a>
</li>
<li>
,
<a href="#!/search/Kaufman%20Howard L./%7B%22type%22:%22author%22%7D">
Howard L. Kaufman
<span>
MD
</span>
</a>
</li>
<li>
and
<a href="#!/search/Miller%20David M./%7B%22type%22:%22author%22%7D">
David M. Miller
<span>
MD, PhD
</span>
</a>
</li>
</ul>
<p class="source" data-once-text="XocsCtrl.citation">
Journal of the American Academy of Dermatology, 2021-10-01, Volume 85, Issue 4, Pages 1038-1040, Copyright © 2021 American Academy of Dermatology, Inc.
</p>
</div>
</div>
<!-- ngIf: ContentCtrl.srctype.toLowerCase() === 'book' && XocsCtrl.hubEid && !ContentCtrl.showPaywall -->
<p ck-reading-mode-toggle="" class="hideprint reading-mode-toggle ng-isolate-scope" data-load-content-function="XocsCtrl.loadAll" data-load-refs-function="ContentCtrl.referenceLoader.loadAllReferences" data-source-type="ContentCtrl.srctype">
<button ck-tooltip="Messages.content_reading_mode_open" class="expand icon icon-expand ng-scope">
<span class="visuallyhidden">
Open reading mode
</span>
</button>
<button ck-tooltip="Messages.content_reading_mode_close" class="icon icon-contract collapse ng-scope" data-tooltip-placement="bottom-left">
<span class="visuallyhidden">
Close reading mode
</span>
</button>
</p>
</header>
<div class="ng-hide" data-once-text="XocsCtrl.preview" ng-show="ContentCtrl.showPaywall">
</div>
<div class="ng-hide" data-once-text="Messages.content_loading_error" ng-show="XocsCtrl.error">
There was an error loading this content. Please refresh the page to try again, or contact us if you continue to experience problems.
</div>
<!-- ngRepeat: item in XocsCtrl.sections -->
<div class="s-content ng-scope early-item" ng-class="{'early-item': $index < 2}" ng-repeat="item in XocsCtrl.sections">
<div data-once-html="item">
<style class="ng-scope">
.c-ckc-abstract{border-bottom:1px solid #d7d7d7;margin-bottom:2em;padding-bottom:1em}.c-ckc-acknowledgment{background-color:#e7e7e7;margin-top:2em;padding:1em}.c-ckc-appendices{border-top:1px solid #d7d7d7;margin-top:2em;padding-top:2em}.c-ckc-article-footnote{margin-top:1em}.c-ckc-author-group{font-weight:700}.c-ckc-bibliography__header{margin-top:1.5em}.c-ckc-bibliography__item{margin:1em 0}.c-ckc-view-in-source-link{margin-right:1em}.c-ckc-cross-reference-external-link{margin-left:1em}.c-ckc-def-list-item{border-bottom:1px dotted #d7d7d7;margin-bottom:1.5em;padding-bottom:.5em}.c-ckc-figure{margin:1.5em 0}.c-ckc-figure__image{height:auto;max-width:100%}.c-ckc-figure__caption,.c-ckc-figure__label,.c-ckc-figure__source{color:#737373;font-size:.875em;margin-bottom:.5em}.c-ckc-figure__label{font-weight:700;text-transform:uppercase}.c-ckc-footnote{color:#737373;font-size:.875em;margin:.5em 0}.c-ckc-formula{margin:1em;text-align:center}.c-ckc-further-reading__header{margin-top:1.5em}.c-ckc-inline-figure{max-width:100%;height:auto}.c-ckc-inline-figure--icon{max-height:2em;width:auto}.c-ckc-journal-head-matter{border-top:1px solid #d7d7d7;margin-top:2em;padding-top:2em}.c-ckc-list{padding-left:1em;list-style:initial;margin-bottom:1.5em;overflow-wrap:break-word}.c-ckc-list__item{list-style:initial}.c-ckc-list__item::marker{color:#e9711c}.c-ckc-list .c-ckc-list{margin-left:1em}.c-ckc-list__item .c-ckc-list__item::marker{color:#969696}.c-ckc-list__item .c-ckc-list__item .c-ckc-list__item{list-style-type:circle}.c-ckc-list__item .c-ckc-list__item .c-ckc-list__item::marker{color:#969696}.c-ckc-list__item .c-ckc-list__item .c-ckc-list__item .c-ckc-list__item{list-style-type:square}.c-ckc-list__item .c-ckc-list__item .c-ckc-list__item .c-ckc-list__item::marker{color:#969696}.c-ckc-list__item-label{float:left;margin-left:-1em}.c-ckc-inline-reference+.c-ckc-inline-reference{margin-left:.25em}.c-ckc-math{display:inline-block;max-width:100%;overflow-x:auto}.c-ckc-math mjx-assistive-mml{padding:0!important;width:1px!important;clip:rect(0 0 0 0)}.c-ckc-section{overflow-wrap:break-word}.c-ckc-section__label{float:left;margin:0 .5em 0 0}.c-ckc-section-title__inline-figure,.c-ckc-section-title__inline-figure--icon{max-height:1em;width:auto}.c-ckc-table{margin:1.5em 0}.c-ckc-table__table{border:1px solid #dcdcdc;border-collapse:collapse}.c-ckc-table__caption,.c-ckc-table__footnote,.c-ckc-table__label,.c-ckc-table__legend{color:#737373;font-size:.875em;margin-bottom:.5em}.c-ckc-table__label{font-weight:700;text-transform:uppercase}.c-ckc-table__superscript-label{color:#007398}.c-ckc-table__overflow{margin:.5em 0;overflow:auto}.c-ckc-table__header-cell{background-color:#ebebeb;font-weight:700;border:1px solid #dcdcdc;padding:.25em;text-align:left;vertical-align:top}.c-ckc-table__body-cell{background-color:#fff;border:1px solid #dcdcdc;padding:.25em;vertical-align:top}.c-ckc-textbox{background-color:#dff8ff;margin:1.5em 0}.c-ckc-textbox__caption,.c-ckc-textbox__label,.c-ckc-textbox__legend,.c-ckc-textbox__source,.c-ckc-textbox__subtitle,.c-ckc-textbox__title{color:#737373;font-size:.875em;margin-bottom:.5em}.c-ckc-textbox__label{font-weight:700;text-transform:uppercase}.c-ckc-textbox__body{padding:1em 1.5em}.c-ckc-textbox__tail{margin-top:1em}.u-ckc-small-cap{font-variant:small-caps}.u-ckc-monospace{font-family:monospace}.u-ckc-superscript{line-height:1}.u-ckc-unstyled-list{list-style-type:none!important}.u-ckc-pull-right{float:right}.u-ckc-clearfix::after{content:"";clear:both;display:table}
</style>
<p class="ng-scope" id="hl0000427">
<i>
To the Editor:
</i>
Immunotherapy has revolutionized the treatment of advanced cutaneous squamous cell carcinoma (advCSCC) not amenable to curative surgery and/or radiotherapy. Phase I/II clinical trials
<button class="j-inline-reference inline-reference u-els-color-linkblue" data-refid="bib1" id="refInSitubib1">
<sup>
1
</sup>
</button>
<sup>
,
</sup>
<button class="j-inline-reference inline-reference u-els-color-linkblue" data-refid="bib2" id="refInSitubib2">
<sup>
2
</sup>
</button>
excluded poor performance status (PS) and immunosuppressed patients. Data on the efficacy of anti-programmed cell death 1 (PD-1) therapy in real-world cohorts are lacking, especially in the advCSCC population generally deemed trial ineligible.
</p>
<p class="ng-scope" id="hl0000434">
We performed an Institutional Review Board-approved retrospective study of patients with advCSCC who received immune checkpoint inhibitors from 2016 to 2020 at Massachusetts General Hospital. Response to immune checkpoint inhibitors was evaluated using Response Evaluation Criteria In Solid Tumors (RECIST) version 1.1. The Kaplan-Meier method was used to estimate overall survival (OS), progression-free survival (PFS), and duration of clinical benefit. In a preplanned exploratory analysis, univariable and multivariable Cox proportional hazards regression were used to model associations between clinicopathologic features and PFS or OS (for details see the Supplementary Methods via Mendeley at
<a class="u-els-color-linkblue" href="https://doi-org.treadwell.idm.oclc.org/10.17632/g769x5dt5r.1" id="hl0000435" target="_blank">
https://doi-org.treadwell.idm.oclc.org/10.17632/g769x5dt5r.1
</a>
).
</p>
<p class="ng-scope" id="hl0000436">
Of the 76 patients that met inclusion criteria (median age, 74 years), 43 patients (57%) had unresectable/locally advCSCC only, and 33 (43%) had distant metastatic disease (
<a ck-scroll-to="" class="u-els-color-linkblue ng-scope" href="tbl1" id="hl0000437" update-fn="scrollToFunc">
Table I
</a>
). Given standard of care guidelines at the time of treatment, 47 patients (62%) received anti–PD-1 therapy as first-line systemic therapy, 17 patients (22%) as second-line, and 12 patients (16%) had ≥2 lines of prior systemic therapy. Nineteen patients (25%) were immunosuppressed, and 26 patients (34%) had an Eastern Cooperative Oncology Group Performance Status ≥2. Additional clinicopathologic characteristics are summarized in
<a ck-scroll-to="" class="u-els-color-linkblue ng-scope" href="tbl1" id="hl0000439" update-fn="scrollToFunc">
Table I
</a>
.
<a id="hl0000009">
</a>
</p>
<div class="table ng-scope" id="tbl1">
<div class="inline-table-label c-content-table__label">
Table I
</div>
<div class="inline-table-caption c-content-table__caption">
Patient characteristics at time of immune checkpoint inhibitor (
<i>
ICI
</i>
) initiation
</div>
<div class="content-overflow">
<table class="c-content-table" id="hl0000014">
<thead>
<tr>
<th align="" class="c-content-table__header-cell" id="hl0000019" scope="col">
Clinical featuresHTML Extraction Pipeline
source(here::here("scripts", "load_packages.R"))
library(reticulate)
## Section 0. Python environment setup
library(reticulate)
py_require(c(
"pandas",
"requests",
"beautifulsoup4",
"lxml",
"python-dotenv",
"selenium",
"webdriver-manager",
"tqdm"
))
## Section 1. Imports and setup
import os
import re
import ast
import time
import json
import shutil
import random
import mimetypes
import subprocess
import urllib.request
from datetime import datetime
from pathlib import Path
from urllib.parse import urlparse, urljoin
import pandas as pd
import requests
from bs4 import BeautifulSoup
from dotenv import load_dotenv
from selenium import webdriver
from selenium.webdriver.chrome.service import Service
from selenium.webdriver.chrome.options import Options
from selenium.webdriver.common.by import By
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.common.exceptions import (
TimeoutException,
ElementClickInterceptedException,
StaleElementReferenceException
)
from webdriver_manager.chrome import ChromeDriverManager
PROJECT_ROOT = Path("/Users/davidmiller/Partners HealthCare Dropbox/David Miller/mLab/Projects/Statistical Methods in Dermatology")
METADATA_DIR = PROJECT_ROOT / "jaad_data" / "metadata"
HTML_ROOT = PROJECT_ROOT / "jaad_data" / "html"
PDF_ROOT = PROJECT_ROOT / "jaad_data" / "pdf"
SUPPLEMENT_ROOT = PROJECT_ROOT / "jaad_data" / "supplements"
load_dotenv()
for path in [METADATA_DIR, HTML_ROOT, PDF_ROOT, SUPPLEMENT_ROOT]:
path.mkdir(parents=True, exist_ok=True)
## Section 2. Helper functions
def get_version(cmd):
"""
Return a version string from a shell command, or None on failure.
"""
try:
result = subprocess.run(cmd, capture_output=True, text=True)
return result.stdout.strip()
except Exception:
return None
def get_major_version(version_string):
"""
Extract major version number from a version string.
"""
if not version_string:
return None
match = re.search(r"(\d+)\.", version_string)
return int(match.group(1)) if match else None
def cleanup_old_chromedriver():
"""
Remove /usr/local/bin/chromedriver if it is incompatible with local Chrome.
Helps avoid driver mismatch errors.
"""
chromedriver_path = Path("/usr/local/bin/chromedriver")
if not chromedriver_path.exists():
print("No chromedriver in /usr/local/bin -> nothing to clean")
return
chrome_version = get_version(
["/Applications/Google Chrome.app/Contents/MacOS/Google Chrome", "--version"]
)
driver_version = get_version([str(chromedriver_path), "--version"])
if chrome_version is None or driver_version is None:
print("Could not determine versions -> skipping cleanup")
return
chrome_major = get_major_version(chrome_version)
driver_major = get_major_version(driver_version)
print(f"Chrome version: {chrome_version}")
print(f"Chromedriver version: {driver_version}")
if chrome_major != driver_major:
print("Removing incompatible chromedriver from /usr/local/bin")
chromedriver_path.unlink()
else:
print("Chromedriver version matches Chrome -> keeping")
def get_driver_cookies_as_requests_session(driver):
"""
Create a requests session that shares Selenium browser cookies.
This lets us download files directly with requests while preserving access.
"""
session = requests.Session()
for cookie in driver.get_cookies():
session.cookies.set(
cookie["name"],
cookie["value"],
domain=cookie.get("domain"),
path=cookie.get("path", "/")
)
session.headers.update({
"User-Agent": driver.execute_script("return navigator.userAgent;"),
"Accept": "application/pdf,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8",
"Accept-Language": "en-US,en;q=0.9",
"Referer": driver.current_url,
})
return session
def safe_filename_piece(x):
"""
Make a filename-safe string.
"""
x = str(x)
keep = []
for ch in x:
if ch.isalnum() or ch in ("-", "_", "."):
keep.append(ch)
else:
keep.append("_")
out = "".join(keep)
while "__" in out:
out = out.replace("__", "_")
return out.strip("_")
def get_extension_from_response(response, fallback=".pdf"):
"""
Infer extension from response headers or final URL.
"""
content_type = (response.headers.get("Content-Type") or "").split(";")[0].strip().lower()
if content_type:
guessed = mimetypes.guess_extension(content_type)
if guessed:
return guessed
final_url = getattr(response, "url", "") or ""
suffix = Path(urlparse(final_url).path).suffix
if suffix:
return suffix.lower()
return fallback
def save_stream_response_to_file(response, save_path):
"""
Save a streamed requests response to disk.
"""
save_path.parent.mkdir(parents=True, exist_ok=True)
with open(save_path, "wb") as f:
shutil.copyfileobj(response.raw, f)
return save_pathHTML Extraction Pipeline
<html class="modern" id="ng-app" lang="en-US" xmlns:ng="http://angularjs.org">
<head>
<style>
@charset "UTF-8";[ng\:cloak],[ng-cloak],[data-ng-cloak],[x-ng-cloak],.ng-cloak,.x-ng-cloak,.ng-hide:not(.ng-hide-animate){display:none !important;}ng\:form{display:block;}.ng-animate-shim{visibility:hidden;}.ng-anchor{position:absolute;}
</style>
<meta content="IE=edge" http-equiv="X-UA-Compatible"/>
<meta charset="utf-8"/>
<meta content="1" name="tdm-reservation"/>
<meta content="https://www-elsevier-com.treadwell.idm.oclc.org/tdm/tdmrep-policy.json" name="tdm-policy"/>
<meta content="width=device-width, initial-scale=1.0, maximum-scale=1.0, user-scalable=no" name="viewport"/>
<title>
Real-world assessment of response to anti-programmed cell death 1 therapy in advanced cutaneous squamous cell carcinoma - ClinicalKey
</title>
<script src="https://js-agent.newrelic.com/nr-spa-1216.min.js">
</script>
<script async="" src="https://cdn.pendo.io/agent/static/b3541d7b-4788-4b73-7811-976020af677d/pendo.js">
</script>
<script type="text/javascript">
;window.NREUM||(NREUM={});NREUM.init={distributed_tracing:{enabled:true},privacy:{cookies_enabled:true},ajax:{deny_list:["bam.nr-data.net"]}};
;NREUM.loader_config={accountID:"1574307",trustKey:"2038175",agentID:"243284150",licenseKey:"94f48af4f8",applicationID:"243284150"}
;NREUM.info={beacon:"bam.nr-data.net",errorBeacon:"bam.nr-data.net",licenseKey:"94f48af4f8",applicationID:"243284150",sa:1}
</path>
<polygon points="29.471 11.883 29.471 15.02 32.609 15.02 32.609 24.435 35.75 24.435 35.75 15.02 38.888 15.02 38.888 11.883 29.471 11.883">
</polygon>
<path d="M15.55139,11.66574c-1.33253,0-2.43215.03784-3.62968,0.118l-0.23665.02446V24.43481H14.8243V19.55419c0,0.01413.47114,0.01413,0.63653,0.01413a4.164,4.164,0,0,0,4.34788-4.00956C19.8087,13.117,18.126,11.66574,15.55139,11.66574ZM14.8243,14.16984l0.65362-.01041c1.06617,0,1.33771.40442,1.33771,1.40753,0,0.71372-.37319,1.54619-2.13379,1.54619H14.8243V14.16984Z">
</path>
<path d="M48,0L0,0.003V35.5574H16.25546L14.7367,40.00025h-5.848v4.44726H39.10989V40.00025H33.25958l-1.518-4.44285H48V0ZM20.25909,40.00025l1.51656-4.44285h4.43907l1.51952,4.44285H20.25909ZM43.5557,31.1109H4.44506V4.44726H43.5557V31.1109Z">
</path>
</symbol>
<symbol id="els-hmds-icon-ppt-2" viewbox="0 0 48 64">
<title>
ppt-2
</title>
<use xlink:href="#icon__ppt-2" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-printer-2" viewbox="0 0 126 128">
<title>
printer-2
</title>
<path d="m97 54h1e1v1e1h-1e1v-1e1zm-6e1 28h52v24h-52v-24zm-1e1 34h72v-44h-72v44zm1e1 -1e2h52v2e1h-52v-2e1zm75 2e1h-13v-3e1h-72v3e1h-13c-7.16 0-13 5.83-13 13v4e1c0 7.17 5.84 13 13 13h5v-1e1h-5c-1.62 0-3-1.37-3-3v-4e1c0-1.63 1.38-3 3-3h98c1.62 0 3 1.37 3 3v4e1c0 1.63-1.38 3-3 3h-5v1e1h5c7.16 0 13-5.83 13-13v-4e1c0-7.17-5.84-13-13-13">
</path>
</symbol>
<symbol id="els-gizmo-icon-publication-set" viewbox="0 0 122 128">
<title>
publication-set
</title>
<path d="m12 57c0-6.08 4.92-11 11-11h17v-2e1h-6c-2.2 0-4 1.8-4 4v6h-7c-3.32 0-6.44 0.78-9.22 2.16 2.44-5.64 7.28-11.86 13.5-17.1 2.34-1.98 5.3-3.06 8.32-3.06h46.4v44.12l8.26-8.26c0.56-0.56 1.14-1.06 1.74-1.54v-44.32h-56.4c-5.38 0-10.62 1.92-14.76 5.4-9.1 7.68-18.84 20.14-18.84 32.1v70.5h41.84l3.12-1e1h-34.96v-49zm97.42 16.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56z">
</path>
</symbol>
<symbol id="els-gizmo-icon-publication-sets" viewbox="0 0 122 128">
<title>
publication-sets
</title>
<path d="m109.42 73.02l-4.18 4.18-6.42-6.42 4.18-4.18c0.64-0.64 1.74-0.66 2.4 0l4.02 4.02c0.64 0.64 0.64 1.74 0 2.4zm-36.16 36.14l-9.32 2.86 2.9-9.3 24.92-24.92 6.42 6.42-24.92 24.94zm43.22-45.62l-4.02-4.02c-2.2-2.2-5.14-3.42-8.28-3.42-3.12 0-6.06 1.22-8.28 3.42l-37.9 37.9-9.26 29.76 29.82-9.18 37.9-37.9c4.58-4.58 4.58-12 0.02-16.56zm-104.48 3.46c0-6.08 4.92-11 11-11h17v-2e1h-6c-2.2 0-4 1.8-4 4v6h-7c-3.32 0-6.44 0.78-9.22 2.16 2.44-5.64 7.28-11.86 13.5-17.1 2.34-1.98 5.3-3.06 8.32-3.06h34.4v46.12l1e1 -1e1v-46.12h-44.4c-5.38 0-10.62 1.92-14.76 5.4-9.1 7.68-18.84 20.14-18.84 32.1v60.5h41.84l3.12-1e1h-34.96v-39zm76-10.88l2.26-2.26c2.2-2.2 4.86-3.82 7.74-4.76v-49.1h-44.4c-5.38 0-10.62 1.92-14.76 5.4-1.64 1.38-3.3 2.94-4.92 4.6h54.08v46.12z">
</path>
</symbol>
<symbol id="els-gizmo-icon-radiology" viewbox="0 0 126 128">
<title>
radiology
</title>
<path d="m48 68.5v18.32c0 5.78-2.04 10.74-6.08 14.7-6.48 6.4-15.98 8.44-19.26 8.4-7.18-0.1-10.66-4.46-10.66-13.34 0-20.48 8.76-51.82 20.08-61.68 4.42-3.86 6.6-2.86 7.32-2.52 2.1 0.96 3.94 3.36 5.4 6.44 2.64-3.02 4.4-6.74 4.98-10.74-1.74-2.08-3.8-3.7-6.22-4.8-4-1.82-10.38-2.58-18.04 4.08-15.14 13.2-23.52 49.26-23.52 69.22 0 14.44 7.68 23.16 20.52 23.34h0.24c5.8 0 17.82-3.02 26.2-11.28 5.92-5.84 9.04-13.38 9.04-21.82v-24.84c-2.72 2.12-3.8 2.7-1e1 6.52zm52.5-41.14c-7.66-6.68-14.04-5.9-18.04-4.08-2.44 1.1-4.48 2.74-6.22 4.82 0.58 4 2.34 7.72 4.98 10.74 1.32-2.8 3.92-6.82 6.96-6.82 1.2 0 3.06 0.56 5.76 2.88 11.3 9.84 20.06 41.2 20.06 61.68 0 8.86-3.48 13.24-10.66 13.34-3.34 0-12.78-2-19.26-8.4-4.04-3.96-6.08-8.92-6.08-14.7v-18.32c-6.18-3.8-7.28-4.38-1e1 -6.52v24.84c0 8.44 3.12 15.98 9.06 21.82 8.38 8.26 20.4 11.28 26.2 11.28h0.24c12.82-0.18 20.5-8.9 20.5-23.34 0-19.96-8.38-56.04-23.5-69.22zm-24.1 30.76l14.22 8.76 5.24-8.52-14.24-8.76c-8.4-5.18-13.62-14.54-13.62-24.42v-19.18h-1e1v19.18c0 9.88-5.22 19.24-13.64 24.42l-14.24 8.76 5.24 8.52 14.22-8.76c5.64-3.48 10.22-8.34 13.4-14 3.2 5.66 7.78 10.52 13.42 14z">
</path>
</symbol>
<symbol id="els-gizmo-icon-rainbow" viewbox="0 0 128 128">
<title>
rainbow
</title>
<path d="m105.76 112h-40.8c-5 0-9.08-4.08-9.08-9.1 0-4.7 2.84-8.12 7.78-9.38l4.06-1.02-0.32-4.2c-0.4-5.38 1-9.98 4.06-13.28 2.98-3.24 7.44-5.02 12.5-5.02 8.06 0 14.9 5.8 16.28 13.8l0.66 3.84 3.88 0.3c7.76 0.6 12.96 5.44 12.96 12.06s-5.38 12-11.98 12zm-63.86-44h-26.06c-3.24 0-5.86-2.74-5.86-6.12 0-3.9 3.32-5.9 6.62-6.16l3.88-0.3 0.66-3.84c0.74-4.4 4.36-7.58 8.62-7.58 2.68 0 5 0.94 6.58 2.62 1.66 1.8 2.42 4.4 2.2 7.5l-0.3 4.16 4.04 1.04c1.64 0.44 3.62 1.56 3.62 4.46 0 2.38-1.76 4.22-4 4.22zm67.3 10.48c-3.24-10.14-12.2-17.4-22.84-18.36-6.36-15.62-16.02-18.12-21.5-18.12-6.72 0-12.22 2.92-16.46 8.58-0.46-4.16-2.08-7.88-4.76-10.76-3.48-3.76-8.4-5.82-13.88-5.82-7.92 0-14.82 5.04-17.52 12.38-7.28 1.96-12.24 8.04-12.24 15.5 0 8.88 7.1 16.12 15.84 16.12h26.06c7.7 0 13.98-6.38 13.98-14.22 0-1.8-0.34-3.52-0.9-5.1 3.34-5.88 7-6.68 9.88-6.68 4.16 0 8.02 3.26 11 9.12-4.54 1.3-8.58 3.7-11.72 7.1-4.12 4.44-6.46 10.36-6.74 16.92-7.18 3.2-11.5 9.72-11.5 17.76 0 10.54 8.54 19.1 19.06 19.1h40.8c12.1 0 21.96-9.88 21.96-22 0-10.64-7.6-19.22-18.52-21.52m-44.34-54.48c-7.22 0-13.94 2.32-19.88 6.54 1.62 1.08 3.14 2.36 4.5 3.82 1 1.08 1.86 2.24 2.64 3.46 3.94-2.46 8.22-3.82 12.74-3.82 10.38 0 19.58 7.14 26.04 18.72 5.08 1.04 9.76 3.18 13.78 6.22-7.64-21.2-22.28-34.94-39.82-34.94m-28.24 2.68c8.12-6.92 17.74-10.68 28.24-10.68 25.52 0 45.8 23.16 51.94 56.48 4.38 1.78 8.18 4.42 11.2 7.7-4.54-43.14-30.16-74.18-63.14-74.18-16.52 0-31.18 7.62-42.28 20.82 2.3-0.66 4.7-1 7.18-1 2.36 0 4.66 0.32 6.86 0.86">
</path>
</symbol>
<symbol id="els-gizmo-icon-rainbow-2" viewbox="0 0 128 128">
<title>
rainbow-2
</title>
<path d="m64 66c-15.44 0-28 15.26-28 34h1e1c0-13.24 8.08-24.02 18-24.02s18 10.78 18 24.02h1e1c0-18.74-12.56-34-28-34m0-18c-25.8 0-46 22.84-46 52h1e1c0-23.56 15.82-42 36-42s36 18.44 36 42h1e1c0-29.16-20.2-52-46-52m0-18c-35.88 0-64 30.74-64 7e1h1e1c0-33.64 23.72-6e1 54-6e1s54 26.36 54 6e1h1e1c0-39.26-28.12-7e1 -64-7e1">
</path>
</symbol>
<symbol id="els-gizmo-icon-rar-file" viewbox="0 0 92 128">
<title>
rar-file
</title>
<path d="m34.01 48l3.03-1e1h0.08l2.84 1e1h-5.95zm-0.46-18l-8.55 26h7.19l1.04-4h7.38l0.98 4h7.41l-8.46-26h-6.99m29.6 12h-6.15v-6h5.8c2.44 0 3.17 1.48 3.17 3 0 2.16-1.63 3-2.82 3zm9.21-4.12c0-4.56-3.14-7.88-6.77-7.88h-14.59v26h6v-8h6.21c2.53 0 2.7 2.2 2.88 4.48 0.08 1.26 0.2 1.52 0.52 3.52h6.39c-0.58-2-0.61-3.94-0.7-5.12-0.23-3-1.3-5.2-3.05-5.98 2.12-0.9 3.11-4.54 3.11-7.02m-65.36-1.88h5.8c2.44 0 3.17 1.48 3.17 3 0 2.16-1.63 3-2.82 3h-6.15v-6zm0 12h6.21c2.53 0 2.7 2.2 2.88 4.48 0.08 1.26 0.2 1.52 0.52 3.52h6.39c-0.58-2-0.61-3.94-0.7-5.12-0.23-3-1.3-5.2-3.05-5.98 2.12-0.9 3.11-4.54 3.11-7.02 0-4.56-3.14-7.88-6.77-7.88h-14.59v26h6v-8m-6-38v1e1h8e1v60.96l-26.93 27.04h-43.07v-42h-1e1v52h57.22l32.78-32.92v-75.08h-9e1m42 9e1h1e1v-2e1h2e1v-1e1h-3e1v3e1">
</path>
</symbol>
<symbol id="icon__rationale" viewbox="0 0 47.47561 47.99999">
<title>
rationale
</title>
<rect height="3.94076" width="3.93866" x="25.56554" y="24.30582">
</rect>
<path d="M27.93008,7.75887a5.97685,5.97685,0,0,0-6.3053,6.30409h3.94076c0-1.57732.78822-3.15155,2.36454-3.15155a2.122,2.122,0,0,1,2.32783,2.54327c-0.28854,2.32673-4.69237,3.08043-4.69237,7.19782v1.28988H29.5042V21.15428c0-2.82861,4.729-3.79544,4.729-7.87942C34.23318,10.1222,31.54779,7.75887,27.93008,7.75887Z">
</path>
<path d="M21.7949,48H17.82614V45.2931c-3.58981.39516-8.15716,0.52118-9.96392-1.03365a2.87839,2.87839,0,0,1-1.03365-2.20511V33.05516H0L7.20973,19.144a17.36781,17.36781,0,0,1,1.3868-7.99784C10.98687,5.24361,18.03077-.74294,29.07133.07538A19.60255,19.60255,0,0,1,43.312,8.056c3.85035,5.18543,5.08433,12.00861,3.4757,19.215-0.66969,3.00083-2.88991,4.8862-4.8496,6.55073A17.489,17.489,0,0,0,39.48752,36.169a14.3032,14.3032,0,0,0-2.57767,5.87034v5.265h-3.971l0.01941-5.69581a18.18831,18.18831,0,0,1,3.39808-7.88041,20.791,20.791,0,0,1,3.01274-2.933c1.62793-1.38139,3.16335-2.68748,3.5445-4.38762,1.73564-7.7717-.55767-12.983-2.7887-15.98487A15.85123,15.85123,0,0,0,28.7785,4.03344c-8.66113-.63959-14.57657,3.84495-16.504,8.60292a15.78078,15.78078,0,0,0-1.05724,6.3397l0.183,0.74081-0.35965.68259L6.637,29.0864h4.16039V41.36094c1.3954,0.35315,5.16614.24333,8.71924-.29063l2.27832-.34245V48Z">
</path>
</symbol>
<symbol id="els-hmds-icon-rationale" viewbox="0 0 47.47561 64">
<title>
rationale
</title>
<use xlink:href="#icon__rationale" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-record" viewbox="0 0 96 128">
<title>
record
</title>
<path d="m48 23.62c-10.26 0-19.9 3.99-27.14 11.24s-11.24 16.89-11.24 27.14 4 19.89 11.24 27.14 16.9 11.24 27.14 11.24c10.26 0 19.9-3.99 27.14-11.24 7.26-7.25 11.24-16.89 11.24-27.14s-4-19.89-11.24-27.14-16.88-11.24-27.14-11.24zm0 86.38c-12.82 0-24.88-4.99-33.94-14.06s-14.06-21.12-14.06-33.94 5-24.87 14.06-33.94 21.12-14.06 33.94-14.06 24.88 4.99 33.94 14.06c9.06 9.06 14.06 21.12 14.06 33.94s-5 24.87-14.06 33.94-21.12 14.06-33.94 14.06">
</path>
</symbol>
<symbol id="icon__redo" viewbox="0 0 50 45.3691">
<title>
redo
</title>
<path d="M22.685,0C29.0709,0,34.38174,2.33729,39.403,7.35627,40.857,8.811,43.2475,11.2204,45.37,13.35983v-8.731H50v16.6664H33.33281V16.66562h8.79643c-2.12713-2.14558-4.5376-4.57309-5.9993-6.03479-4.151-4.15024-8.2974-6.002-13.44492-6.002A18.05634,18.05634,0,1,0,40.55509,24.99842h4.684A22.679,22.679,0,1,1,22.685,0Z">
</path>
</symbol>
<symbol id="els-hmds-icon-redo" viewbox="0 0 50 64">
<title>
redo
</title>
<use xlink:href="#icon__redo" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="icon__reduce" viewbox="0 0 48 48.00779">
<title>
reduce
</title>
<polygon points="48 3.356 44.64 0 29.696 14.942 29.696 1.215 24.95 1.215 24.95 23.058 46.795 23.058 46.795 18.311 33.045 18.311 48 3.356">
</polygon>
<polygon points="1.203 29.708 14.942 29.708 0 44.65 3.358 48.008 18.3 33.066 18.3 46.805 23.05 46.805 23.05 24.958 1.203 24.958 1.203 29.708">
</polygon>
</symbol>
<symbol id="els-hmds-icon-reduce" viewbox="0 0 48 64">
<title>
reduce
</title>
<use xlink:href="#icon__reduce" xmlns:xlink="http://www.w3.org/1999/xlink">
</use>
</symbol>
<symbol id="els-gizmo-icon-refresh" viewbox="0 0 112 128">
<title>
refresh
</title>
<path d="m74 6e1h36v-36h-1e1v18.86c-4.58-4.62-9.75-9.83-12.89-12.97-10.84-10.84-22.32-15.89-36.11-15.89-27.02 0-49 21.98-49 49s21.98 49 49 49c25.33 0 46.2-19.32 48.72-44h-10.09c-2.46 19.14-18.82 34-38.63 34-21.5 0-39-17.5-39-39s17.5-39 39-39c11.12 0 20.07 4 29.04 12.96 3.16 3.16 8.36 8.41 12.96 13.04h-19v1e1">
</path>
</symbol>
<symbol id="els-gizmo-icon-remove-document" viewbox="0 0 92 128">
<title>
remove-document
</title>
<path d="m29 4e1h34v1e1h-34v-1e1zm14 6e1h1e1v-2e1h2e1v-1e1h-3e1v3e1m38-19.04l-26.93 27.04h-43.07v-88h7e1v60.96zm-8e1 -70.96v108h57.22l32.78-32.92v-75.08h-9e1">
</path>
</symbol>
<symbol id="els-gizmo-icon-repeat" viewbox="0 0 111 128">
<title>
repeat
</title>
<path d="m102.24 42.91-7.16 7.16c2.12 3.29 3.38 6.96 3.38 11.16 0 11.58-9.42 20.77-21 20.77h-44.2l13.38-13.13-7.08-6.96-25.44 25.51 25.44 25.49 7.08-7.29-13.4-13.62h44.22c17.1 0 31-13.67 31-30.77 0-6.96-2.34-13.14-6.22-18.32m-89.78 18.32c0-11.58 9.42-21.23 21-21.23h44.22l-13.38 13.61 7.08 7.18 25.44-25.39-25.44-25.43-7.08 6.85 13.4 13.18h-44.24c-17.1 0-31 14.14-31 31.23 0 6.97 2.34 13.51 6.24 18.69l7.16-7.22c-2.14-3.29-3.4-7.26-3.4-11.47">
</path>
</symbol>
<symbol id="els-gizmo-icon-replay" viewbox="0 0 108 128">
<title>
replay
</title>Natural Language Processing
Deterministic
Code-First/Rule-based extraction
Example: p\\s*[<=>]\\s*0\\.\\d+
Key properties
• Highly reproducible
• Fast runtime
• Transparent logic
Deep Learning
Language Model Interpretation
Example: ChatGPT
Key properties
• Flexible across formats
• Higher computational cost
• Less transparent
Key idea
Differences often involve tradeoffs in:
Reproducibility
Transparency
Scalability
Runtime
Deterministic NLP
######################################
# Read and Process JAAD HTML
######################################
# Load Packages -----------------------------------------------------------
source(here::here("scripts", "load_packages.R"))
# Load Functions ----------------------------------------------------------
source(here::here("scripts", "functions", "functions_read_html.R"))
# Load and Prepare HTML Files ---------------------------------------------
volume_path <- file.path(here(), "jaad_data", "html")
html_files <- dir_ls(volume_path, recurse = TRUE, glob = "*.html")
# Step 1: Basic file metadata ---------------------------------------------
html_file_metadata <- tibble(
file = html_files,
size_kb = round(file_info(html_files)$size / 1024, 1),
file_name = path_file(html_files),
article_id = str_extract(path_file(html_files), "(?<!-)[A-Z0-9]+(?=\\.html$)")
)
###############################
## Filter for target article
#target_article_id <- "0190962219308680"
#html_file_metadata <- html_file_metadata |>
# filter(article_id == target_article_id)
###############################
# Step 2: Read HTML safely once -------------------------------------------
# Add modification time so cache invalidates when file changes
# html_file_metadata <- html_file_metadata %>% mutate(mod_time = file_info(file)$modification_time)
# returns NULL on failure; faster/smaller than storing error objects
read_html_possibly <- possibly(xml2::read_html, otherwise = NULL)
html_loaded <- html_file_metadata %>%
mutate(html_page = map(file, read_html_possibly)) %>%
filter(!map_lgl(html_page, is.null)) # keep only successes
# Step 3: Extract title and main text using html_page ---------------------
# filter for aritcle to inspect
#html_loaded <- html_loaded |> filter(article_id == "0190962215020058")
text_extracted <- html_loaded |>
mutate(
title = map_chr(html_page, extract_jaad_title),
text_full = map_chr(html_page, extract_jaad_text_r),
figure_legend_text = map_chr(html_page, extract_figure_legend_text),
text = map_chr(html_page, extract_text_before_discussion)
)
# Step 4: Clean and add abstract, authors, citation -----------------------
text_annotated <- text_extracted |>
mutate(
title = str_remove_all(title, "Download PDF"),
abstract_text = map_chr(html_page, extract_abstract_text),
main_text_only = pmap_chr(
list(text, abstract_text, figure_legend_text),
function(main_text, abstract, fig_legend) {
temp <- main_text
if (!is.na(abstract) && abstract != "" && str_detect(temp, fixed(abstract))) {
temp <- str_remove(temp, fixed(abstract))
}
if (!is.na(fig_legend) && fig_legend != "" && str_detect(temp, fixed(fig_legend))) {
temp <- str_remove(temp, fixed(fig_legend))
}
str_squish(temp)
}
),
metadata = map(html_page, safely(extract_jaad_metadata_r)),
authors = map_chr(metadata, ~ .x$result$authors %||% ""),
citation = map_chr(metadata, ~ .x$result$citation %||% "")
)
# Step 5: Extract volume/issue/pages --------------------------------------
vol_issues <- text_annotated %>%
mutate(
volume = str_extract(citation, "Volume\\s+\\d+") %>% str_extract("\\d+") %>% as.integer(),
issue = str_extract(citation, "Issue\\s+\\d+") %>% str_extract("\\d+") %>% as.integer(),
page_range = str_extract(citation, "Pages?\\s+[S]?[\\d]+[-–][S]?[\\d]+"),
page_start_chr = str_match(page_range, "Pages?\\s+([A-Za-z]*\\d+)")[,2],
page_start_num = suppressWarnings(as.integer(str_extract(page_start_chr, "\\d+"))),
page_end = str_extract(page_range, "(?<=[-–])[\\d]+") %>% as.integer()
)
#----------------------------------------
# Filter to desired articles using TOC-derived identifiers
#----------------------------------------
toc <- open_recent_file(
directory = "jaad_data/toc",
ext = ".csv",
contains = "jaad_articles"
)
normalize_title <- function(x) {
x %>%
stringr::str_to_lower() %>%
stringr::str_replace_all("&", "and") %>%
stringr::str_replace_all("[^a-z0-9]+", "") %>%
stringr::str_squish()
}
extract_start_page <- function(page_range) {
stringr::str_match(page_range, "Pages?\\s+([A-Za-z]*\\d+)")[, 2]
}
toc <- toc %>%
mutate(
pii_from_doi_path = stringr::str_extract(
doi,
"(?<=/science/article/pii/)[A-Z0-9]+"
),
article_pii = dplyr::coalesce(article_id, pii_from_doi_path),
article_pii = stringr::str_remove(article_pii, "^S"),
start_page = dplyr::coalesce(start_page, extract_start_page(page_range)),
title_norm = normalize_title(title),
fallback_id = stringr::str_glue(
"v{volume}_i{issue}_p{start_page}_{stringr::str_sub(title_norm, 1, 80)}"
),
unique_article_id = dplyr::if_else(
!is.na(article_pii) & article_pii != "",
paste0("pii_", article_pii),
fallback_id
)
)
desired_articles <- c("Research article", "Short communication")
toc_filtered <- toc %>%
filter(article_type %in% desired_articles) %>%
mutate(
has_issue_metadata = !is.na(volume) & !is.na(issue),
has_start_page = !is.na(start_page)
) %>%
arrange(
unique_article_id,
desc(has_issue_metadata),
desc(has_start_page)
) %>%
distinct(unique_article_id, .keep_all = TRUE) %>%
select(-has_issue_metadata, -has_start_page)
#----------------------------------------
# Use TOC to retain desired article types
#----------------------------------------
normalize_title <- function(x) {
x %>%
stringr::str_to_lower() %>%
stringr::str_replace_all("&", "and") %>%
stringr::str_replace_all("[^a-z0-9]+", "") %>%
stringr::str_squish()
}
extract_start_page <- function(page_range) {
stringr::str_match(page_range, "Pages?\\s+([A-Za-z]*\\d+)")[, 2]
}
vol_issues_id <- vol_issues %>%
mutate(
article_pii = dplyr::na_if(article_id, ""),
article_pii = stringr::str_remove(article_pii, "^S"),
start_page = dplyr::coalesce(page_start_chr, extract_start_page(page_range)),
title_norm = normalize_title(title),
fallback_id = stringr::str_glue(
"v{volume}_i{issue}_p{start_page}_{stringr::str_sub(title_norm, 1, 80)}"
),
unique_article_id = dplyr::if_else(
!is.na(article_pii),
paste0("pii_", article_pii),
fallback_id
)
)
vol_issues_unique_article_id_dupes <- vol_issues_id %>%
janitor::get_dupes(unique_article_id)
nrow(vol_issues_unique_article_id_dupes)
nrow(vol_issues_id)
vol_issues_id_toc <- vol_issues_id %>%
left_join(
toc_filtered %>% select(unique_article_id, article_type),
by = join_by(unique_article_id)
)
vol_issues_id_toc_desired <- vol_issues_id_toc %>%
filter(article_type %in% desired_articles)
vol_issues_id_toc_desired %>%
count(unique_article_id, sort = TRUE) %>%
filter(n > 1)
vol_issues_id_toc_all <- vol_issues_id %>%
left_join(
toc %>% select(unique_article_id, article_type),
by = join_by(unique_article_id)
)
table(is.na(vol_issues_id_toc_all$article_type))
#----------------------------------------
types_of_articles <- vol_issues_id_toc_desired %>%
mutate(
text_lower = str_to_lower(str_trim(text)),
title_lower = str_to_lower(title),
abstract_lower = str_to_lower(abstract_text),
full_text_lower = paste(title_lower, abstract_lower, text_lower, sep = " "),
headings_lower = map(
html_page,
~ html_elements(.x, "h2") |> html_text2() |> str_to_lower() |> str_squish()
),
is_to_the_editor_letter = str_detect(text_lower, "^to the editor[:punct:]*"),
is_likely_editorial = str_detect(
text_lower,
"in this issue.*?(jaad|academy of dermatology)|this month in jaad"
),
has_materials_and_methods =
map_lgl(headings_lower, ~ any(str_detect(.x, "^materials and methods$|^methods$"))) |
str_detect(text_lower, "\\b(materials and methods|methods)\\b"),
is_systematic_review_or_meta_analysis =
str_detect(title_lower, "systematic review|meta[- ]analysis") |
str_detect(
full_text_lower,
paste(
"we (conducted|performed|undertook) (a )?(systematic review|meta[- ]analysis)",
"prisma(-statement)?",
"cochrane review",
"according to prisma",
sep = "|"
)
),
is_delphi_or_consensus_study = str_detect(
full_text_lower,
paste(
"delphi (survey|process|method|exercise)",
"consensus (meeting|process|building exercise|statement|was reached|was achieved)",
"core outcome set\\b|\\bcos\\b",
"cosmin",
"comet initiative",
"ideom",
"grappa",
"ppacman",
sep = "|"
)
),
is_game_changer_editorial = str_detect(title, fixed("JAAD Game Changers:", ignore_case = TRUE)),
is_letter_from_editor = str_detect(title, fixed("Letter from the Editor", ignore_case = TRUE)),
is_jaad_international_column = str_detect(title, fixed("This month in JAAD International", ignore_case = TRUE)),
is_jaad_reviews_column = str_detect(title, fixed("This month in JAAD Reviews", ignore_case = TRUE)),
is_jaad_case_reports_column = str_detect(title, fixed("This month in JAAD Case Reports", ignore_case = TRUE)),
is_jaad_monthly_column = str_detect(title, fixed("This month in JAAD", ignore_case = TRUE)),
is_beyond_jaad = str_detect(title_lower, "^beyond jaad\\b"),
is_jaad_cme_title_pattern = str_detect(title, regex("part\\s*[ivx]+\\s*[:\\-]", ignore_case = TRUE)),
is_nonresearch_article_base = detect_nonresearch_type(title)
) |>
group_by(volume, issue) |>
arrange(page_start_num, .by_group = TRUE) %>%
mutate(is_one_of_first_two_articles = row_number() <= 2) |>
ungroup() |>
mutate(
is_jaad_cme = is_jaad_cme_title_pattern & is_one_of_first_two_articles,
is_nonresearch_article = (
is_nonresearch_article_base |
is_game_changer_editorial |
is_likely_editorial |
is_letter_from_editor |
is_jaad_international_column |
is_jaad_reviews_column |
is_jaad_case_reports_column |
is_jaad_monthly_column |
is_beyond_jaad |
is_jaad_cme
),
is_patient_characteristics_table = map2_lgl(
main_text_only,
file,
detect_patient_char_table
)
)
types_of_articles_meta <- types_of_articles |>
left_join(
html_file_metadata,
by = join_by(file, size_kb, file_name,
article_id)) |>
arrange(volume, issue, page_start_num)Benchmarking the Deterministic Pipeline
Across 56 manually reviewed articles and 1839 total analyte instances, the deterministic pipeline demonstrated:
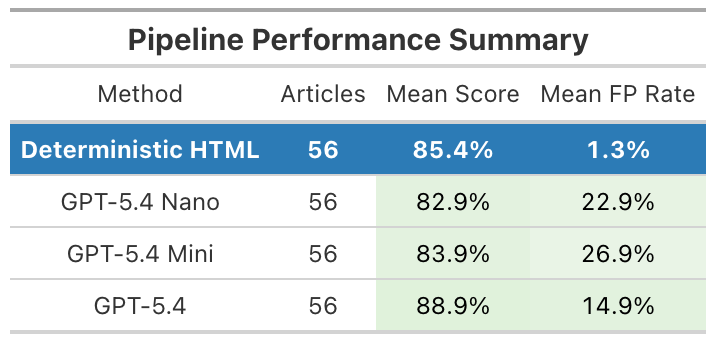
Important
Designed to prioritize specificity over sensitivity
JAAD Statistical Evidence Dataset
We assembled a large corpus of JAAD articles spanning Volumes 74–94 (2016–2026).
Across this interval, we retrieved 11,962 HTML files.
After restricting to Research Articles and Short Communications, the analytic dataset included 4,721 studies.
This provides the empirical foundation for evaluating how statistical evidence is structured within the literature.
Use of P-values in JAAD Research Articles
Among 4,721 studies,
3,242 reported at least one p-value.
This corresponds to 69% of the literature.
Key idea
P-values are a dominant component of statistical evidence in this corpus.
Reported P-Values Per Study
Reported P-Values Per Study
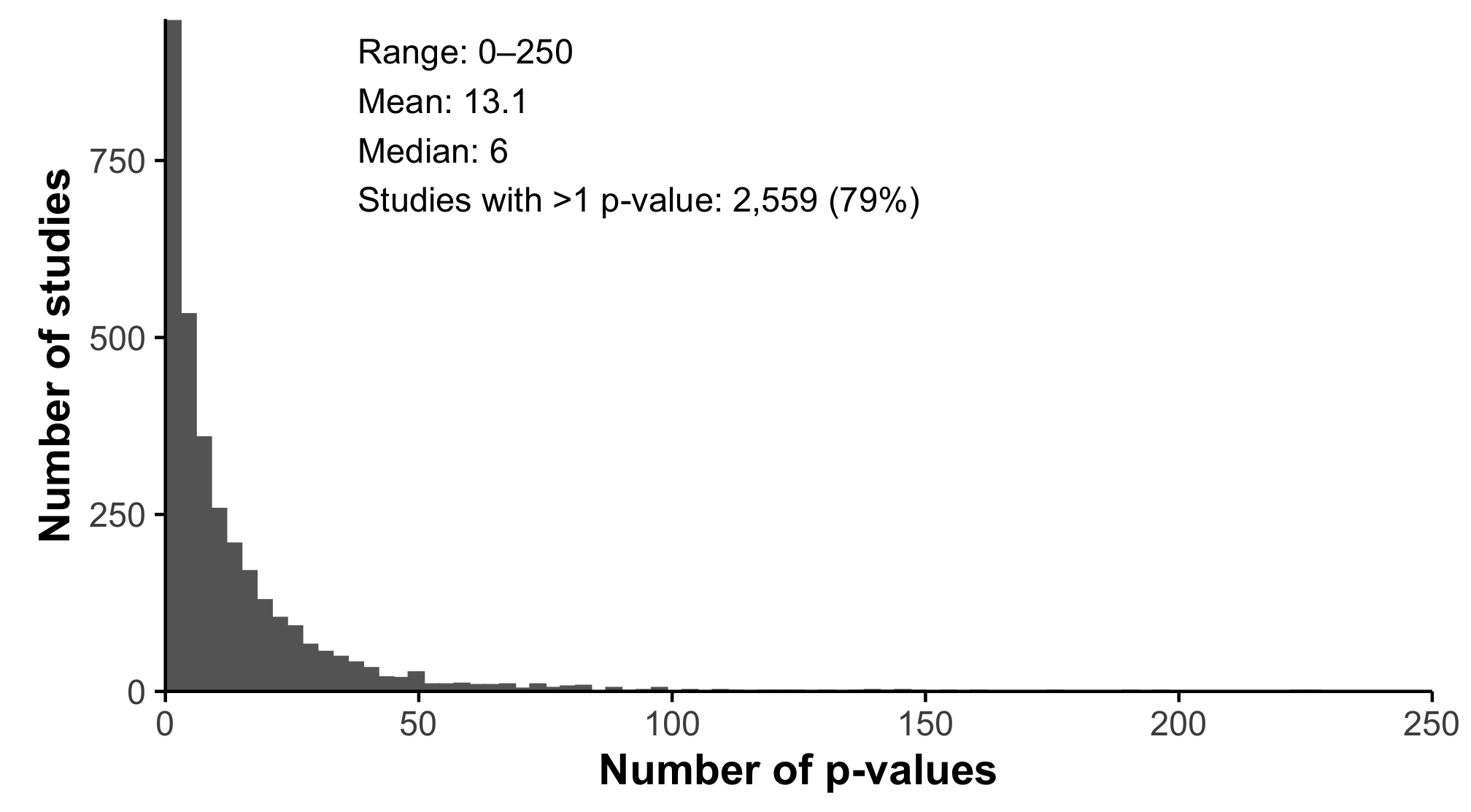
Analytic Spaces in the Real World
Large analytic spaces are a natural feature of clinical research.
Multiple analyses often arise from reasonable scientific questions.
However, as the number of statistical tests increases, the probability of false positive findings also increases.
Statistical safeguards can help preserve interpretability.
Tip
One important safeguard is adjustment for Multiple Hypothesis Testing.
Adjustment for Multiple Hypothesis Testing
Studies with >1 P-value
Adjustment for Multiple Hypothesis Testing
Studies with >1 P-value
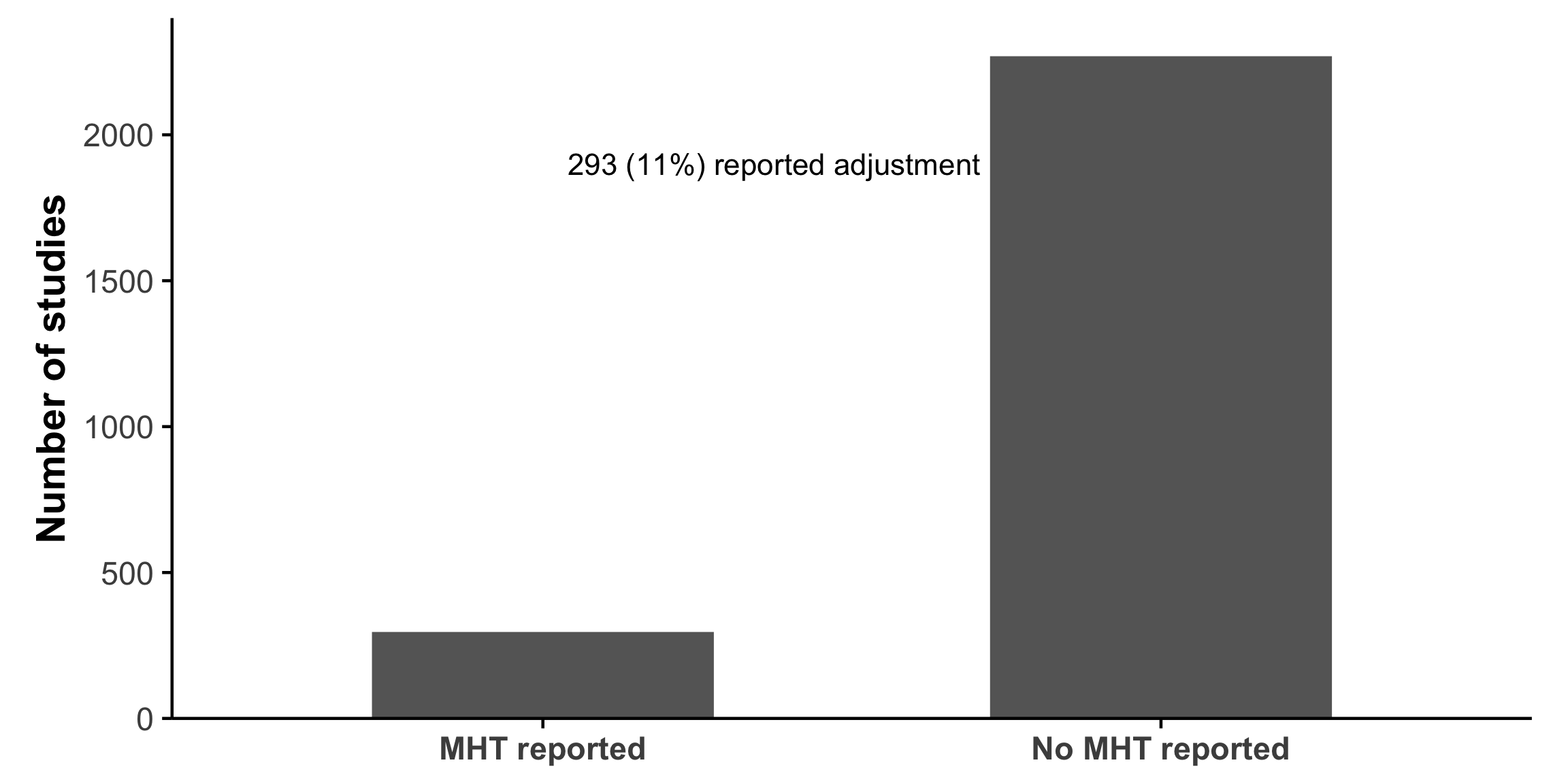
Primary Explicit Multiple-Testing Method
One Primary Method per Article
Primary Explicit Multiple-Testing Method
One Primary Method per Article
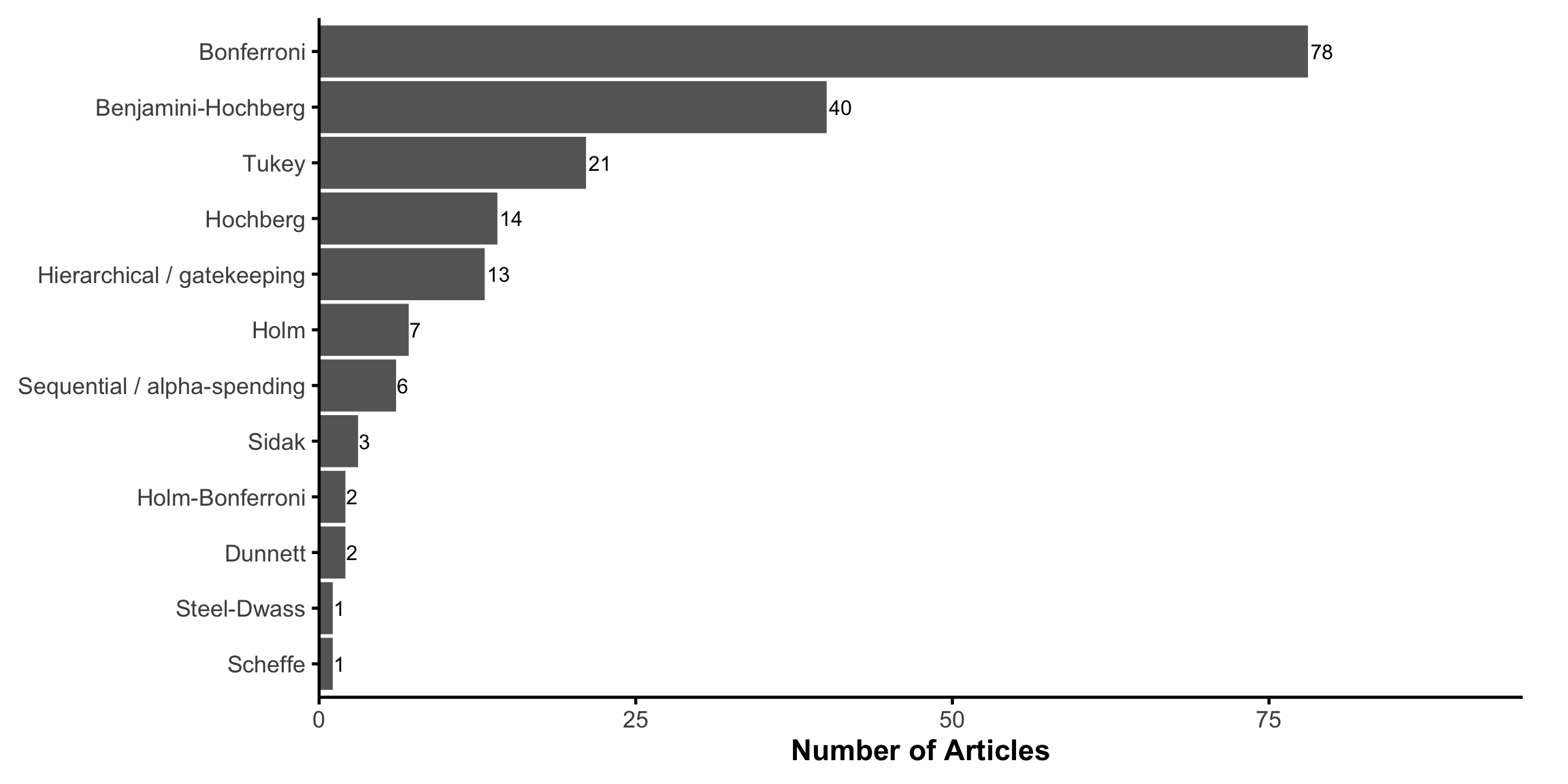
Distribution of Reported P-values
Articles: 4,721 | Total p-values: 42,210
Distribution of Reported P-values
Articles: 4,721 | Total p-values: 42,210
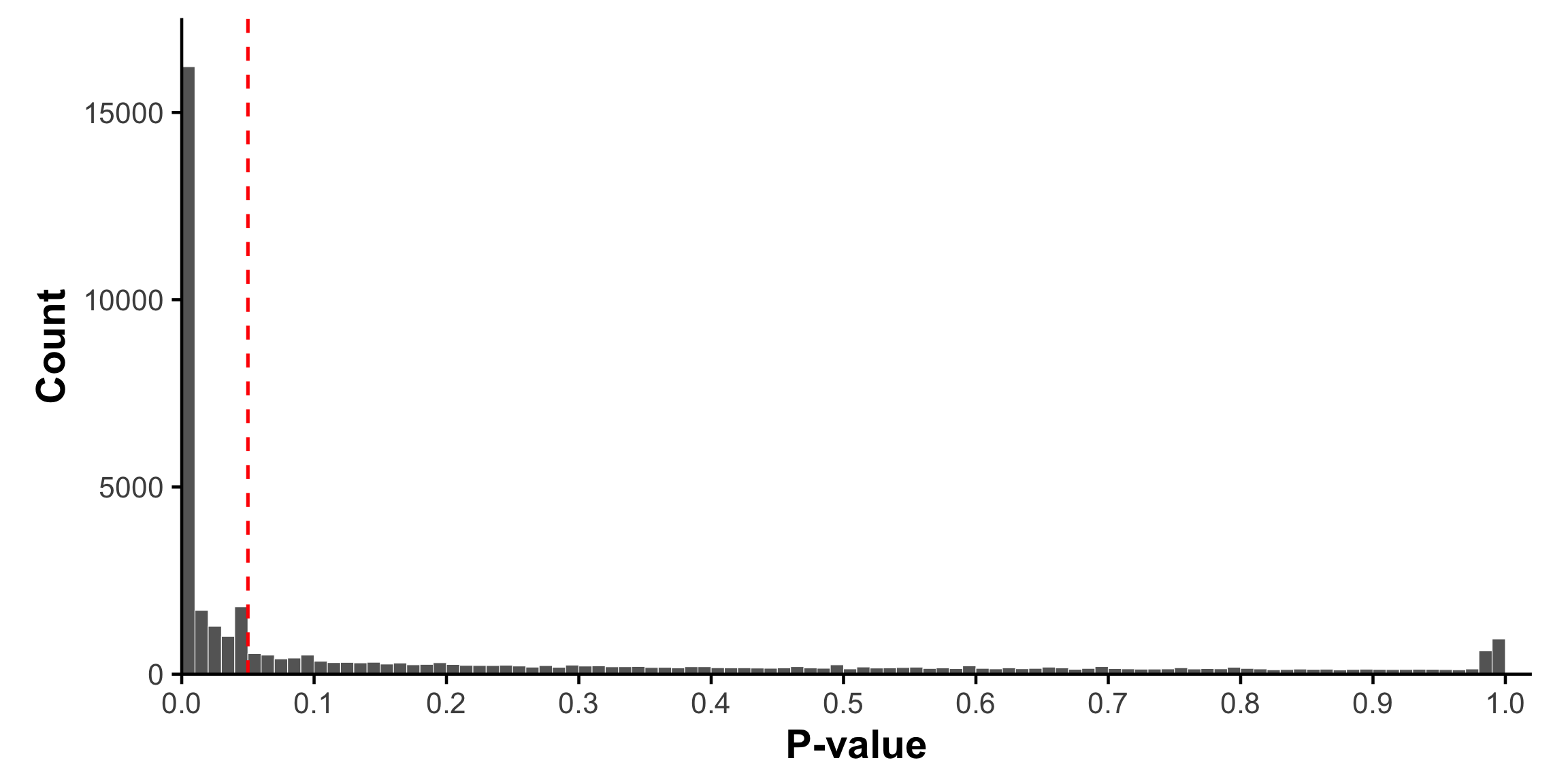
Average Number of P-Values Per Study
Nominal vs FDR vs Bonferroni thresholds
Average Number of P-Values Per Study
Nominal vs FDR vs Bonferroni thresholds
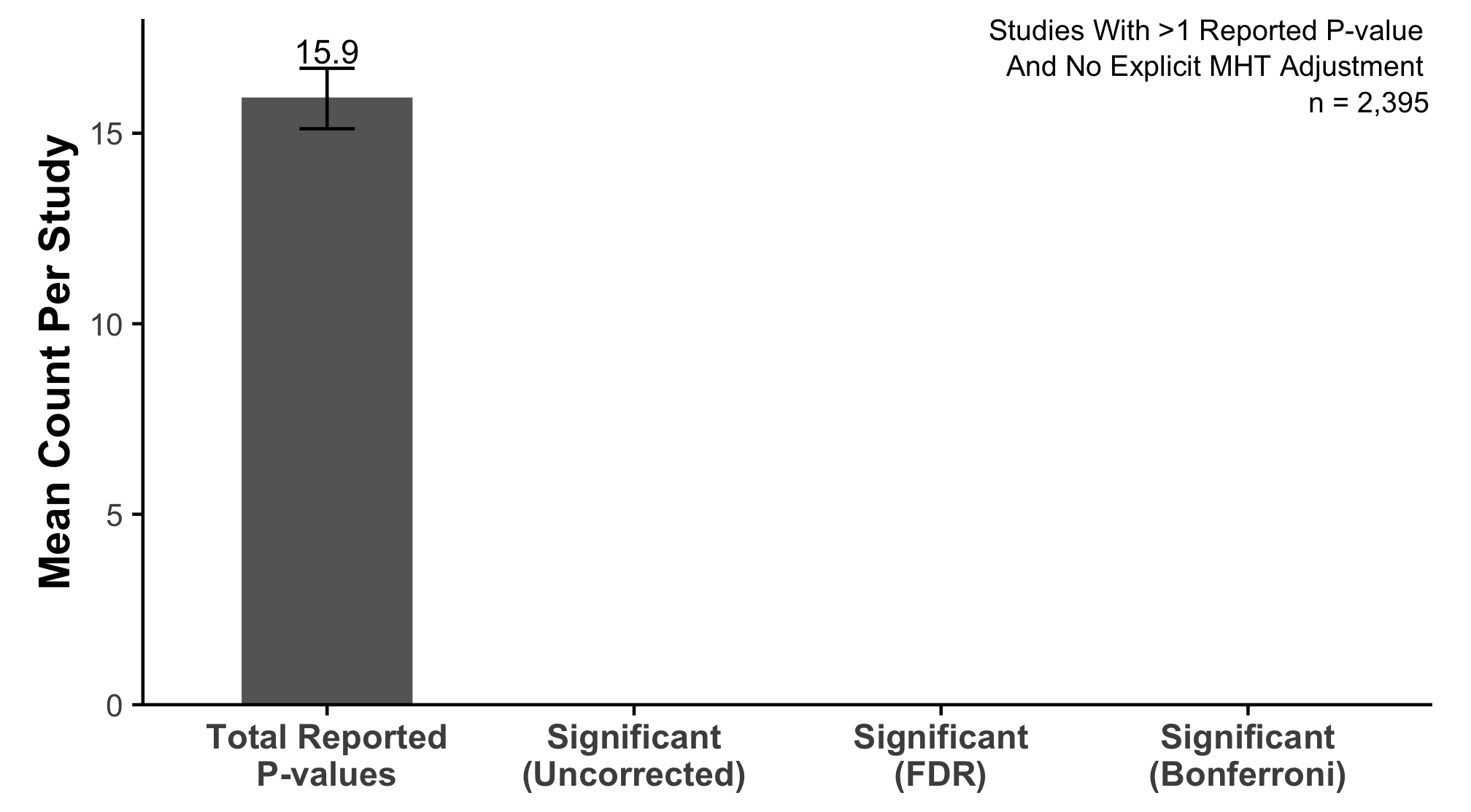
Average Number of P-Values Per Study
Number of P-Values < 0.05
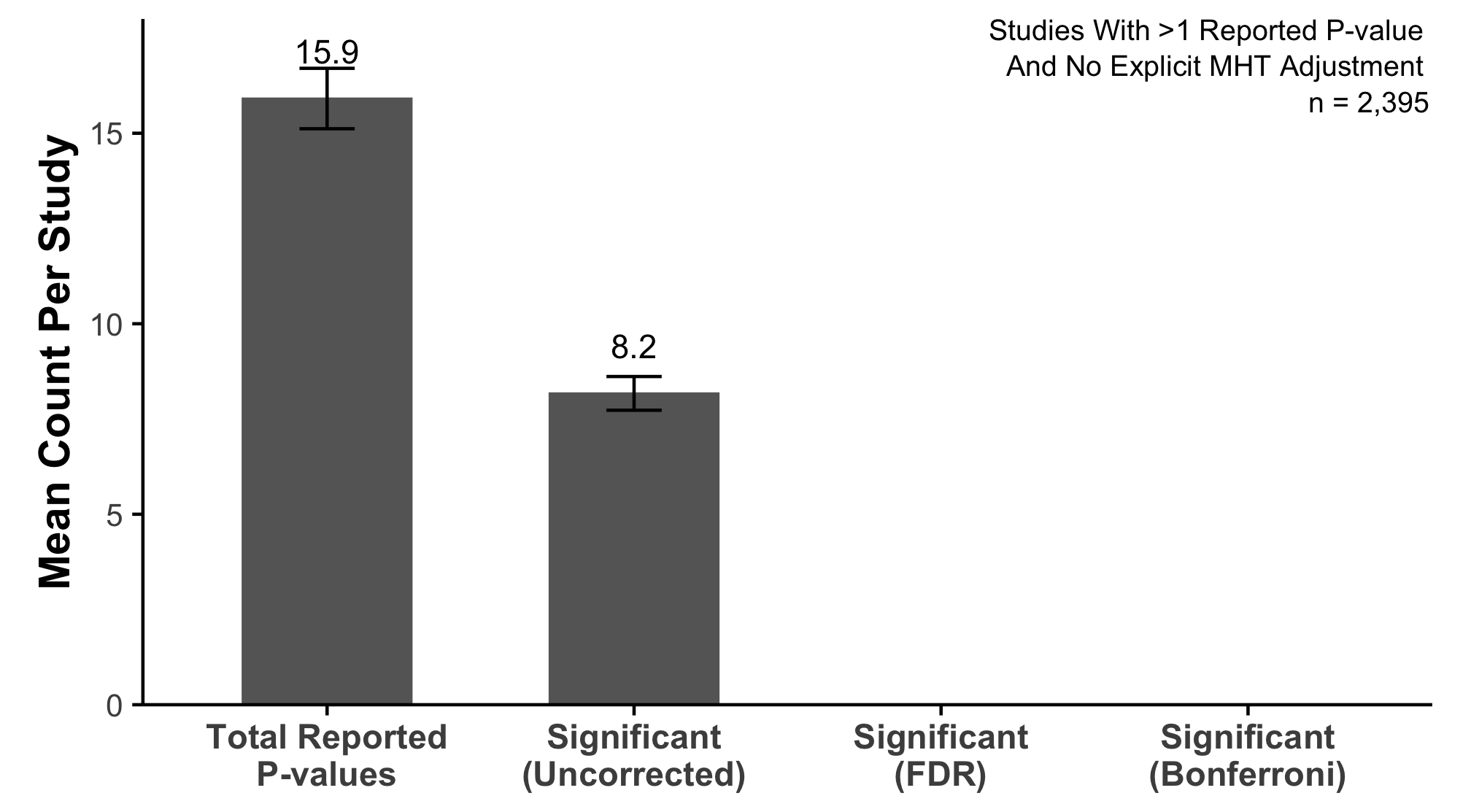
How Many Results Remain Significant?
Nominal vs FDR vs Bonferroni thresholds
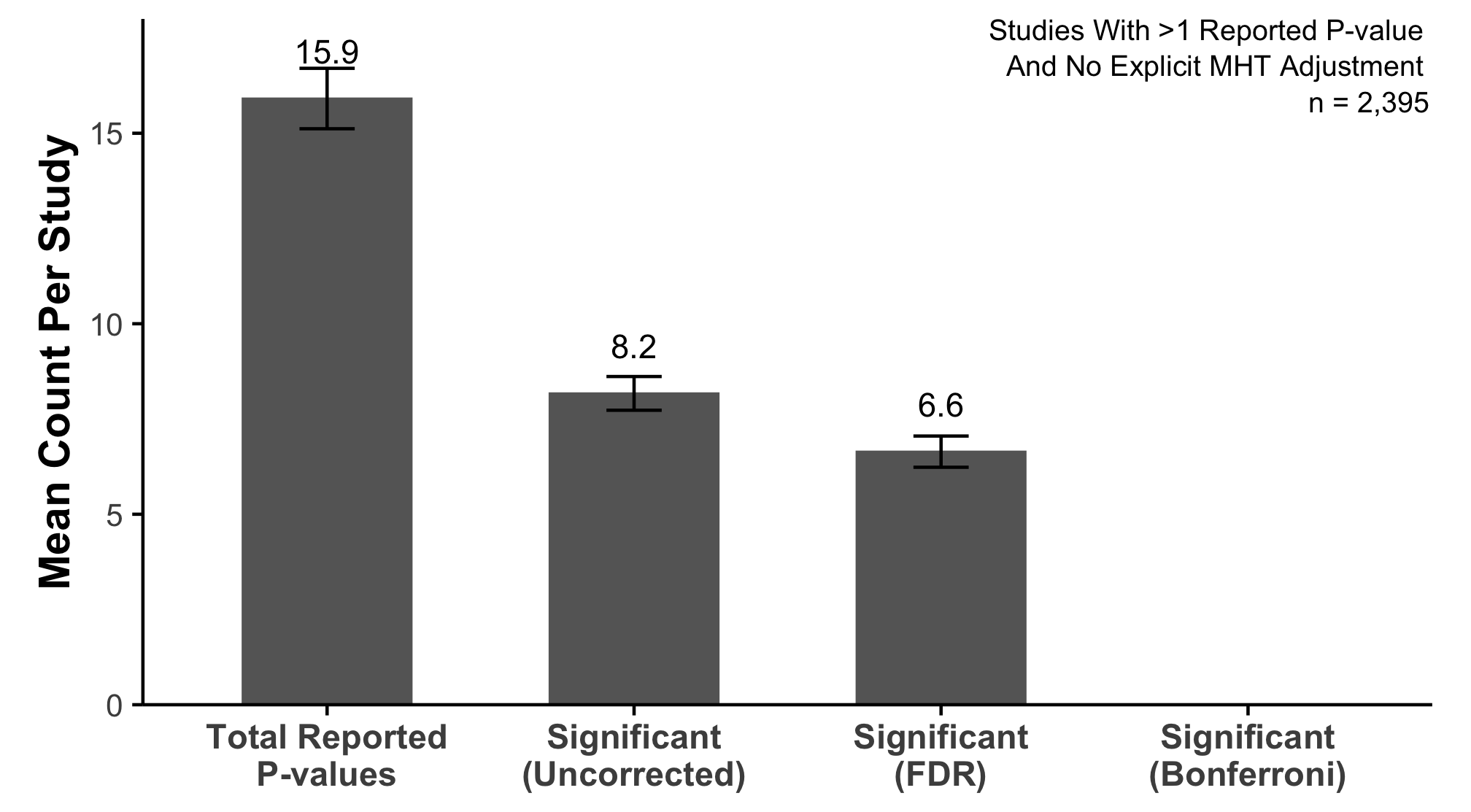
How Many Results Remain Significant?
Nominal vs FDR vs Bonferroni thresholds
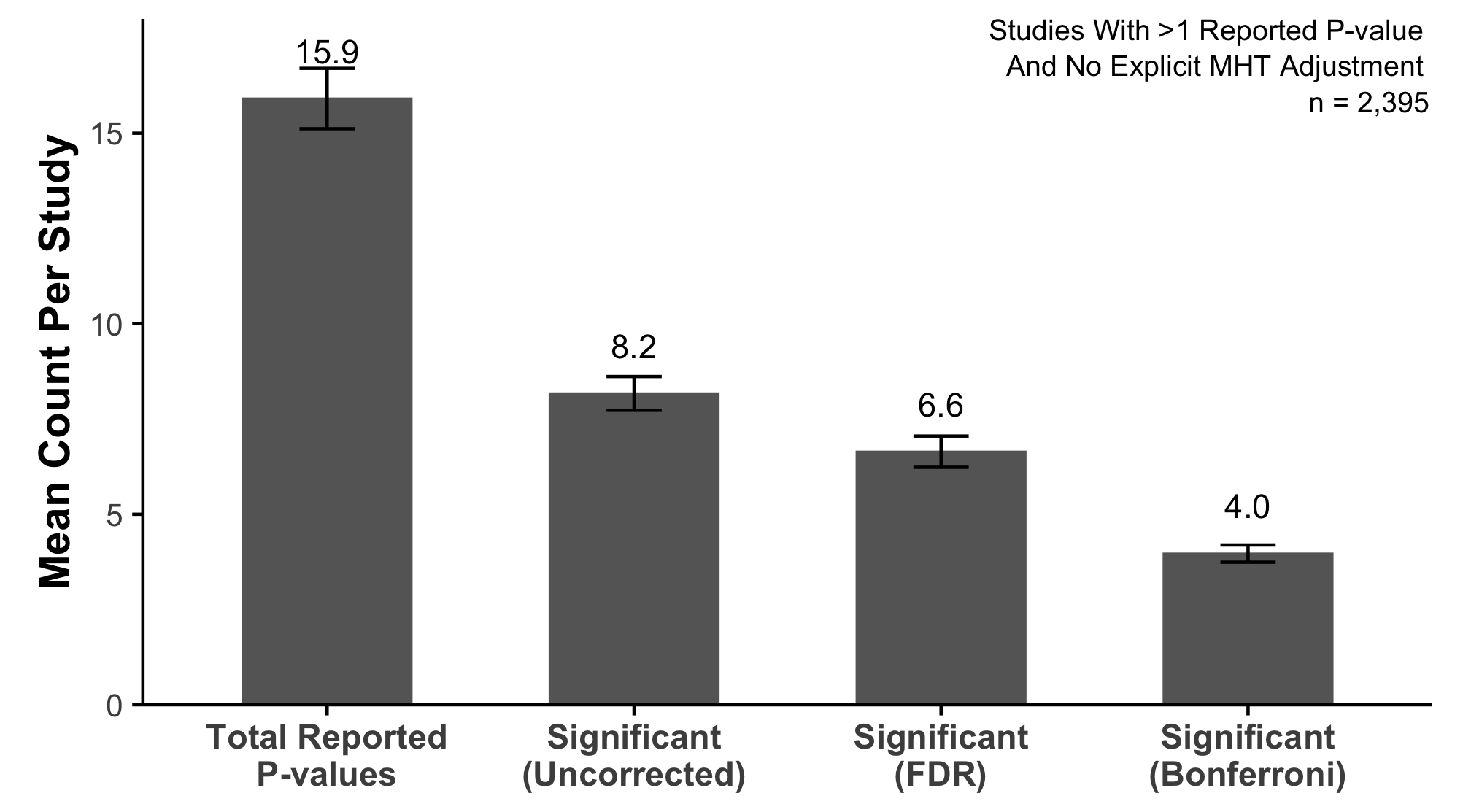
Significant Results Depend on the Threshold Used
In studies with more than one reported p-value and no explicit multiplicity adjustment, the mean number of significant results falls substantially under Bonferroni correction
Using this more conservative threshold, nearly half of nominally significant p-values would no longer be interpreted as statistically significant
Key idea
This does not mean those findings are false
It means their interpretation depends on the inferential framework being applied
What Is Family-Wise Error Rate?
Family-wise error rate is the probability of getting at least one false positive across all statistical tests in a study
If each test uses α = 0.05, that risk increases as the number of tests increases
So even if every individual p-value is judged using the usual 0.05 threshold, the study-level chance of at least one false positive may be much higher than 5%
Important
The more hypotheses tested in a study, the harder it is to interpret any single “significant” result in isolation.
Estimated Family-Wise Type I Error Rate
If studies controlled family-wise error near 5%

Estimated Family-Wise Type I Error Rate
If studies controlled family-wise error near 5%

Estimated Family-Wise Type I Error Rate
Studies Reporting P-values Without Explicit Multiple Testing
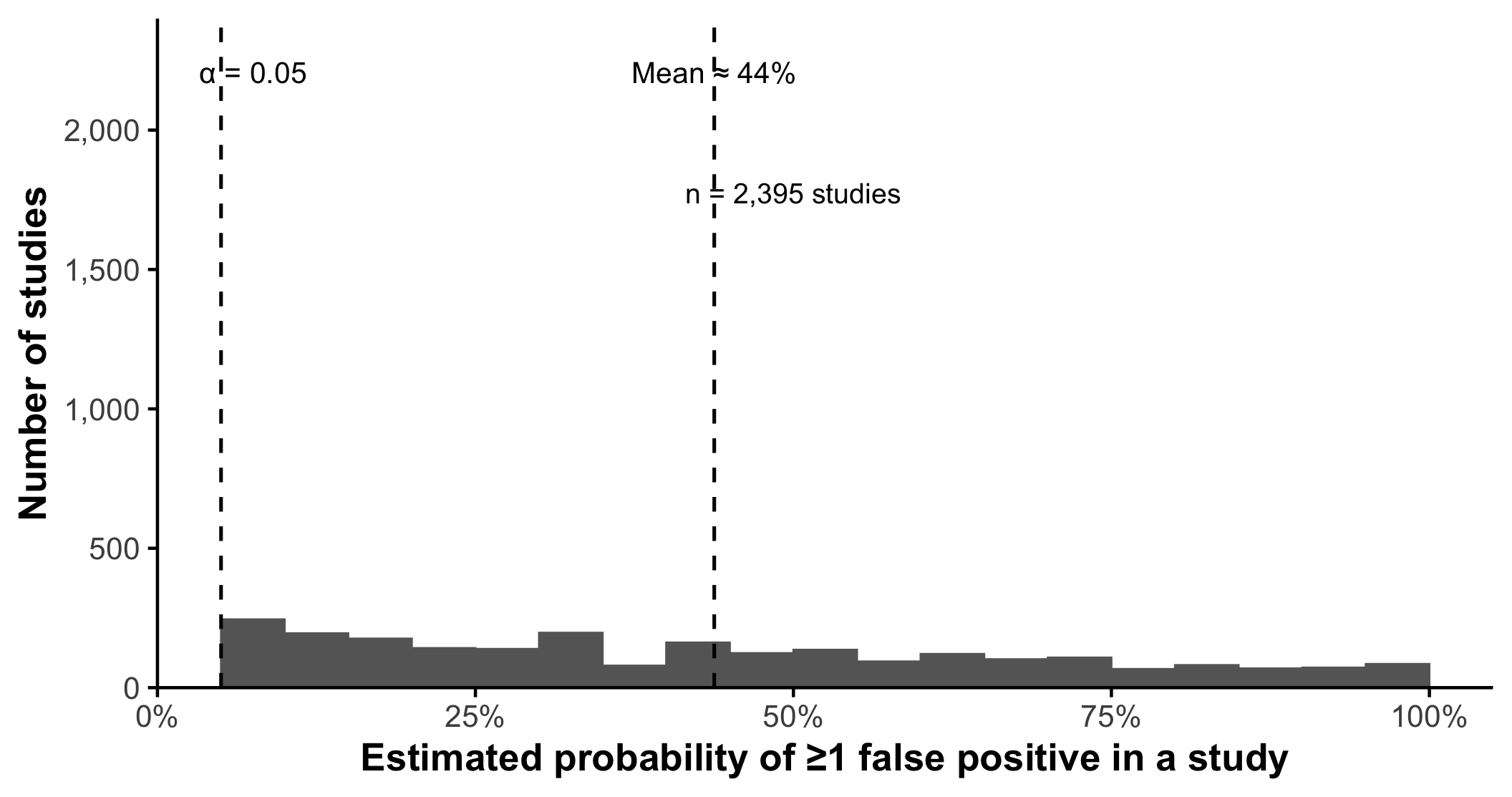
The Standard Varies By Context
In FDA trials and high-impact journal submissions, family-wise error is controlled to 5%
In the literature you read every day — the average estimated false positive risk per study may be as high as ~40%
Important
The evidence shaping routine clinical decisions may reflect a different standard
So how does this happen? It’s not usually deliberate — it’s structural.
What You See In A Paper May Not Be The Whole Story
When you read a paper, you are often seeing the analysis that worked — not all the analyses that were tried
Hypothesis generation ≠ hypothesis testing
• Noticing a pattern in your data and then testing it in the same data is not the same as predicting it in advance
• Because the statistical test assumes you had no idea what you were going to find
Exploratory findings presented as confirmatory evidence can overstate the strength of that evidence
Tip
Preregistration helps clarify which analyses were planned — and which emerged after looking at the data
What is Preregistration?
Document study hypotheses
before examining the data
Specify outcomes, predictors,
and analytic approach a priori
Creates a transparent record
of analytic intent
Example registry:
clinicaltrials.gov
Open Science Framework (OSF)
Tip
Key idea
Preregistration distinguishes
confirmatory analyses
from exploratory analyses
Example Preregistration (OSF) From Our Study
Example Preregistration (OSF) From Our Study
PROJECT DESCRIPTION
Statistical Methods in Clinical and Biomedical Research is an ongoing meta-research project designed to characterize statistical reporting practices in the clinical and translational literature. The initial phase focuses on original research articles published in the Journal of the American Academy of Dermatology (JAAD) between 2016 and March 2026. The methodological framework is designed to be extensible to additional journals and biomedical disciplines in future phases.
The primary objective is to describe patterns in statistical reporting, including:
- prevalence and distribution of reported p-values
- use of multiple comparison adjustments
- adoption of Bayesian statistical approaches
- reporting of preregistration
- analytic structure of reported statistical evidence
To operationalize these measurements at scale, we developed a reproducible two-stage text extraction framework.
1. Deterministic extraction pipeline
A rule-based natural language processing pipeline implemented in R and Python parses article HTML to identify candidate statistical reporting features.
2. LLM-based extraction layer
A secondary extraction approach uses large language models to evaluate content that may be difficult to capture deterministically, including information embedded in tables, figures, and supplementary materials.
Both pipelines are benchmarked against a manually validated gold standard subset of articles.
The deterministic pipeline serves as the primary extraction engine, while the LLM layer functions as a structured sensitivity analysis evaluating whether layout-dependent or supplement-based reporting materially alters article-level conclusions.
The project is intended as a descriptive audit of methodological reporting practices rather than an evaluation of the scientific validity of individual articles.
---------------------------------------------------------------------
FOREKNOWLEDGE OF DATA
This study uses publicly available published articles as the unit of analysis. The data corpus (JAAD articles 2016–2026) existed prior to preregistration and is accessible to the investigators.
A subset of articles was reviewed during development of the extraction pipeline in order to design reproducible text-processing methods and define operational criteria for identifying statistical reporting features.
Pilot work was limited to methodological development and validation of extraction procedures and was not used to finalize hypothesis thresholds or analytic decision rules.
The preregistered hypotheses, variable definitions, and analysis plan were specified prior to execution of the full corpus-wide extraction and inferential analyses.
To reduce risk of unintended analytic flexibility:
- hypotheses were defined prior to full-scale data extraction
- transformation rules for p-values were prespecified
- deterministic and LLM extraction pipelines are applied uniformly across all articles
- sensitivity analyses are prespecified rather than data-driven
- exploratory analyses will be clearly labeled
- all code and intermediate datasets will be publicly shared
---------------------------------------------------------------------
STUDY DESIGN
This study is a computational meta-research analysis of statistical reporting practices in published biomedical literature.
The unit of analysis is the individual research article.
No human subjects are involved.
No experimental intervention is performed.
Study type: Descriptive study
Causal interpretation: No causal relationship inferred
---------------------------------------------------------------------
SAMPLING PLAN
Inclusion criteria:
- Original research articles
- Published in JAAD between 2016 and early 2026
- Machine-readable full text available in HTML or PDF format
Exclusion criteria:
- Editorials
- Commentaries
- Letters
- Case reports
- Systematic reviews
- Meta-analyses
- Consensus statements
- Delphi studies
These exclusions are applied because such article types do not primarily report original statistical analyses.
Sample size:
Based on journal structure, the projected corpus includes approximately 3,000 to 6,000 original research articles.
Because supplementary materials frequently contain large statistical tables, the total number of extracted p-values is expected to substantially exceed 50,000.
All eligible articles will be included.
---------------------------------------------------------------------
VARIABLES AND INDICES
Derived variables characterize statistical reporting patterns at both the article level and p-value level.
Text-reported p-values will be standardized to numeric form using prespecified rules:
P < .05 -> 0.05
P ≤ .01 -> 0.01
P < .001 -> 0.001
P-values will be grouped into prespecified intervals for distributional analyses, including bins near conventional thresholds (e.g., 0.04–0.05 and 0.05–0.06).
Article-level binary indicators will be created for:
- presence of preregistration statements
- presence of Bayesian statistical analyses
- presence of multiple testing correction procedures
- presence of confidence intervals
Multiplicity-adjusted significance counts will be computed using:
Benjamini–Hochberg false discovery rate
Bonferroni correction
Extraction pipeline performance metrics will include:
Precision
Recall
F1 score
All transformations will be applied consistently across deterministic and LLM extraction pipelines.
---------------------------------------------------------------------
DATA COLLECTION PROCEDURES
Articles are retrieved using automated scripts that capture HTML content and associated metadata.
Supplementary materials are downloaded when available in PDF format and linked to the parent article record.
Two independent extraction pipelines identify statistical reporting features.
Deterministic pipeline identifies:
- p-values
- confidence intervals
- multiple testing procedures
- preregistration statements
- Bayesian analyses
LLM pipeline identifies the same features with emphasis on:
- dense tables
- figure captions
- supplement-only reporting
A subset of articles is manually reviewed to estimate extraction accuracy and characterize measurement error.
---------------------------------------------------------------------
HYPOTHESES
Hypothesis 1
Fewer than 5% of original research articles use Bayesian statistical approaches.
Hypothesis 2
Among p-values between 0.04 and 0.06, more than 50% fall below 0.05.
Hypothesis 3
The distribution of reported p-values between 0 and 0.1 deviates from a uniform distribution.
Hypothesis 4
Among studies reporting more than one p-value, fewer than 10% explicitly report a multiple testing correction procedure.
Hypothesis 5
Fewer than 10% of original research articles report preregistration.
Hypothesis 6
Application of standard multiple testing correction procedures reduces the number of statistically significant p-values within studies reporting multiple inferential tests.
---------------------------------------------------------------------
STATISTICAL MODELS
Analyses are conducted at two levels:
1. article-level prevalence estimation
2. p-value-level distributional analysis
Hypotheses 1, 4, and 5 evaluate prevalence of article-level characteristics using one-sample binomial models.
Hypothesis 2 evaluates asymmetry of p-values near 0.05 using a binomial model restricted to the interval 0.04–0.06.
Hypothesis 3 evaluates deviation of the p-value distribution from uniformity using a chi-squared goodness-of-fit test with bin width 0.01.
Hypothesis 6 evaluates the impact of multiplicity correction procedures by comparing counts of statistically significant p-values before and after Benjamini–Hochberg and Bonferroni adjustment using paired Wilcoxon signed-rank tests.
Bayesian models estimate posterior distributions for article-level prevalence parameters using weakly informative priors centered on prespecified reference values.
Extraction pipeline performance relative to manual gold standard annotation will be summarized using precision, recall, and F1 score.
Comparisons between deterministic and LLM extraction pipelines are descriptive and intended to characterize measurement error rather than to test causal hypotheses.
All analyses are prespecified and applied uniformly across the corpus.
---------------------------------------------------------------------
TRANSFORMATIONS
Text-reported p-values will be standardized to numeric form using prespecified rules.
Inequality expressions will be converted to conservative boundary values.
P-values will be grouped into prespecified bins for distributional analyses.
Multiplicity-adjusted p-values will be computed using Benjamini–Hochberg and Bonferroni procedures.
All transformation rules will be applied identically across deterministic and LLM extraction pipelines.
---------------------------------------------------------------------
INFERENCE CRITERIA
Frequentist inference will use one-sided tests for directional hypotheses (Hypotheses 1, 2, 4, 5) and two-sided tests for distributional deviation (Hypothesis 3).
Nominal alpha = 0.05 will be reported for transparency and comparability with the biomedical literature.
Interpretation will emphasize estimation and uncertainty rather than binary significance thresholds.
Bayesian analyses will summarize posterior medians and 95% credible intervals.
Robustness of conclusions will be evaluated using sensitivity analyses including false discovery rate and Bonferroni procedures where applicable.
Exploratory analyses will be clearly labeled.
---------------------------------------------------------------------
DATA INCLUSION AND EXCLUSION
All eligible original research articles published in JAAD between 2016 and early 2026 will be included.
Articles will be excluded only if they do not meet prespecified inclusion criteria.
No exclusions will be performed based on statistical results.
Articles lacking machine-readable full text or accessible supplementary material will be flagged but retained when possible.
No outlier removal procedures will be applied.
---------------------------------------------------------------------
MISSING DATA
Missing data primarily arise from:
- unavailable supplementary materials
- non-machine-readable formatting
- absence of reported statistical quantities within an article
Missing statistical features will not result in exclusion of otherwise eligible articles.
Analyses will use all available extracted information.
Patterns of missingness will be described descriptively.
---------------------------------------------------------------------
OTHER PLANNED ANALYSES
Exploratory analyses may evaluate variation in statistical reporting patterns across:
- publication year
- article structure
- presence of supplementary materials
- density of reported statistical tests
Sensitivity analyses will evaluate robustness of conclusions to differences between deterministic and LLM extraction pipelines.
Exploratory analyses will be clearly labeled.
---------------------------------------------------------------------
OPEN SCIENCE PRACTICES
All code, metadata, and derived datasets will be shared to facilitate reproducibility.
The project is designed to allow extension to additional journals and disciplines using the same extraction framework.Reporting of Preregistration
Across the Curated JAAD Research Corpus
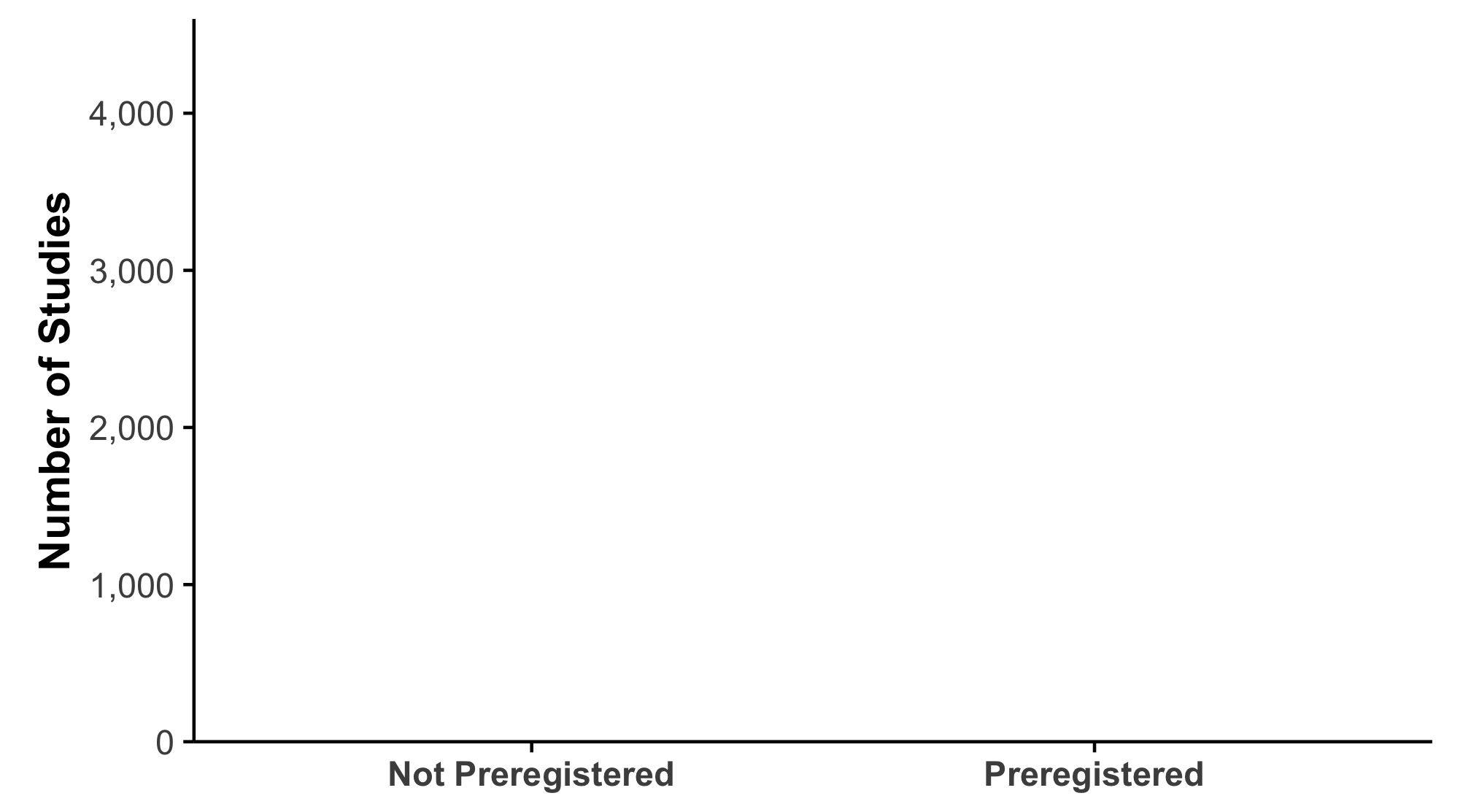
Reporting of Preregistration
Across the Curated JAAD Research Corpus
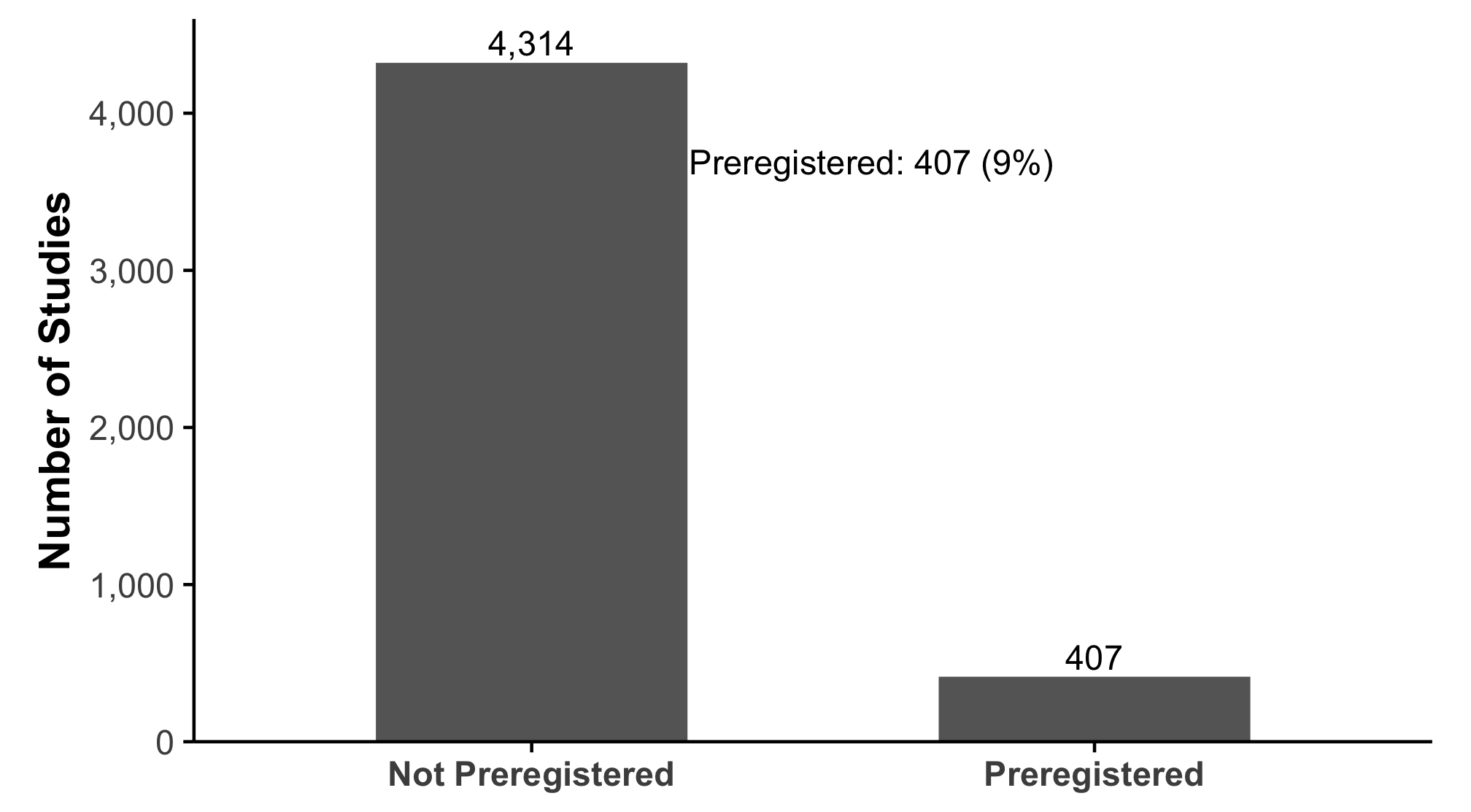
Primary Pre-Registration Platform
Among Pre-Registered Studies

Selective P-value Reporting
Selective reporting can distort how statistical evidence appears in the literature.
Preregistration may help by clarifying which analyses were planned in advance.
Tip
If selective reporting is present, p-values may cluster just below conventional significance thresholds.
A Closer Look At The Threshold
We’ve seen the overall distribution — heavily skewed toward small p-values
But there’s a more specific question:
Are p-values clustering just below 0.05 — right at the significance threshold?
If selective reporting is occurring, we would expect to see more values just below 0.05 than just above it

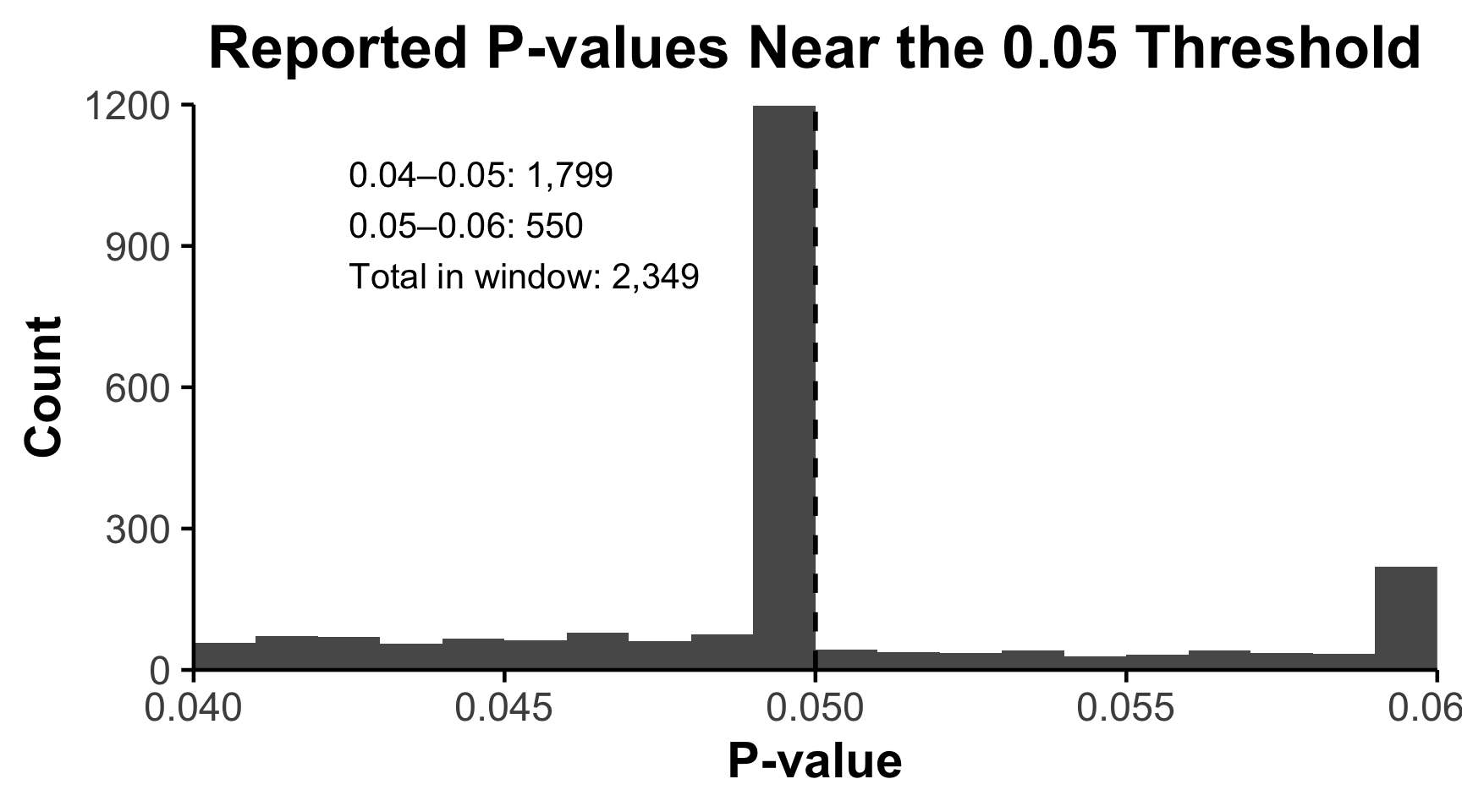
Why Does This Pattern Matter?
Crossing 0.05 Has Often Been Used To Define “Statistical Significance”
A Small Change In P-value Can Change How A Result Is Labeled And Interpreted

If Dichotomizing Evidence at p = 0.05 Creates Instability…
How else can we quantify evidence?
How can we incorporate prior knowledge?
How can we avoid binary decision thresholds?
Alternative Frameworks Express Evidence Continuously
Bayesian Inference Updates Belief Using Data
Start With An Initial Belief
Observe New Data
Update Belief Based On Evidence
Bayes’ theorem
\[ p(\theta \mid D) \;=\; \frac{p(D \mid \theta)\, p(\theta)}{p(D)} \]
Estimand: Is the coin fair?
Alternate Hypothesis: The coin is biased (P(Heads) ≠ 0.5)
Null Hypothesis: The coin is fair (P(Heads) = 0.5)

Experiment: Flip the coin 6 times
Model: Binomial(n = 6, p)
Decision rule: reject H₀ if p-value < 0.05
Frequentist Framework: P(Data | Hypothesis)

Frequentist Framework: P(Data | Hypothesis)

Null hypothesis: H₀: p = 0.5
Likelihood of observed data: P(6 heads | p = 0.5) = 0.56 = 0.0156
p-value: P(X ≥ 6 | p = 0.5) = 0.0156
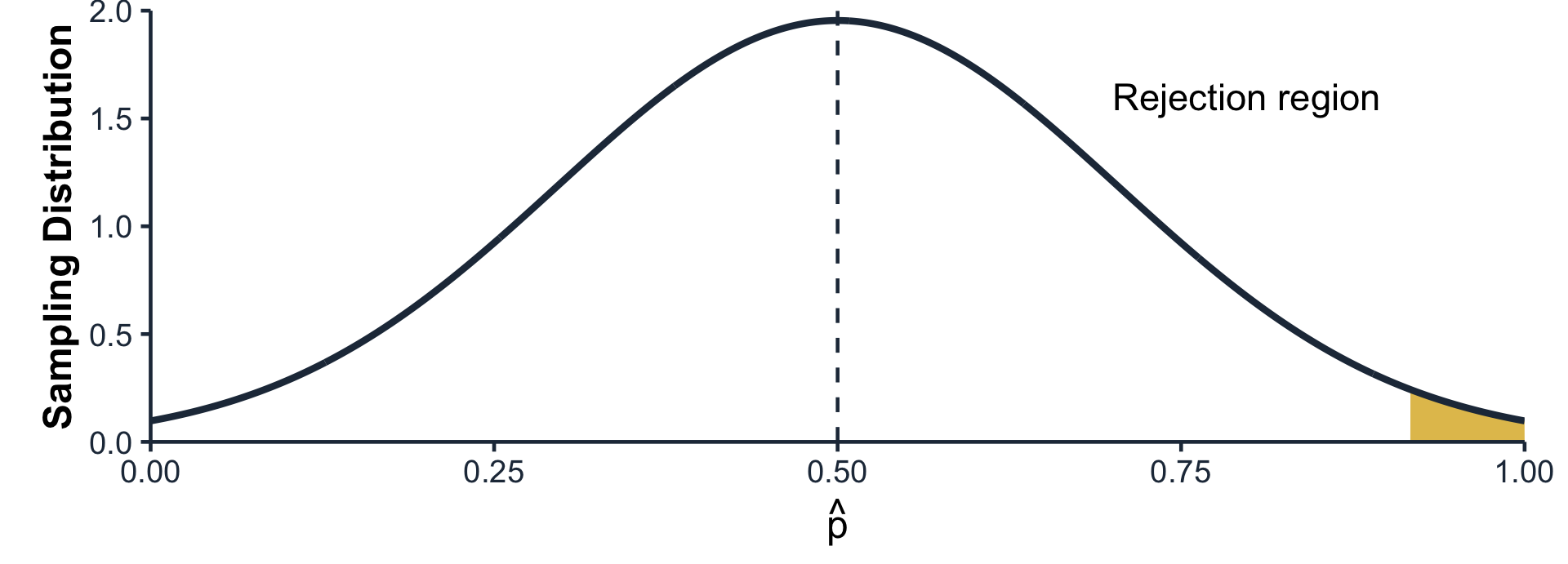
Bayesian Framework: P(Hypothesis | Data)

Prior: Beta(1, 1) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(7, 1)
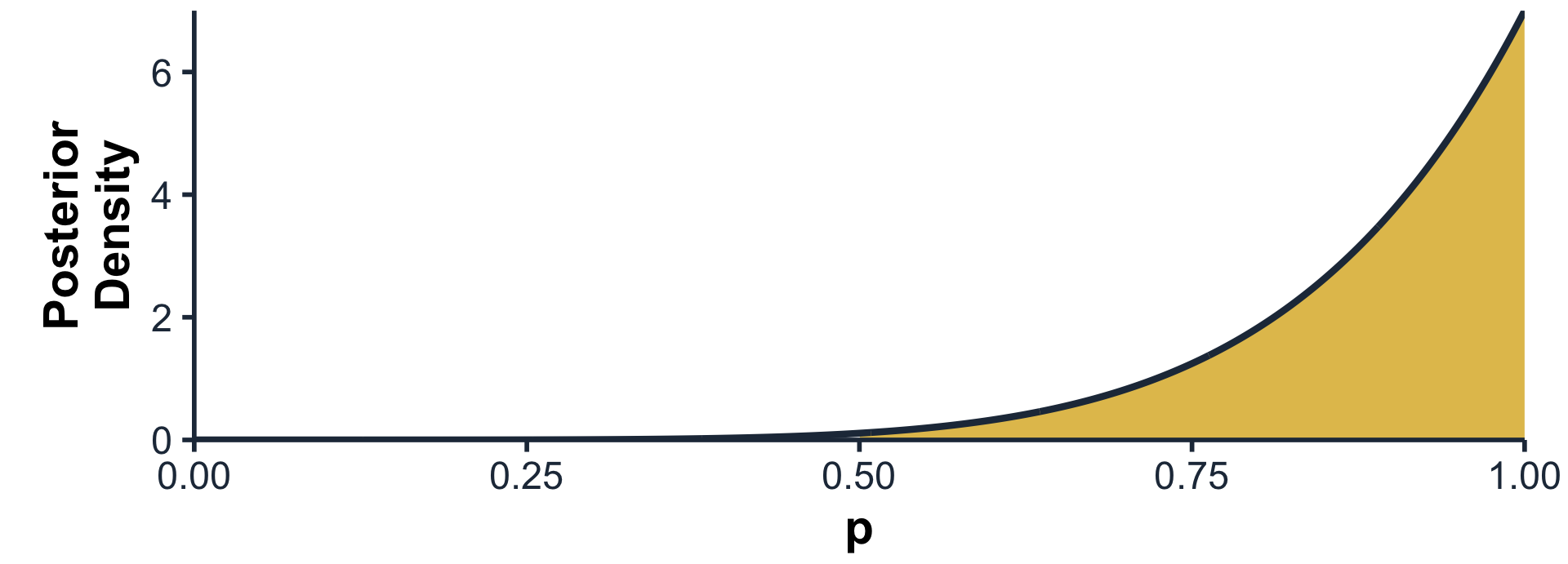
Bayesian Framework: P(Hypothesis | Data)

Prior: Beta(1, 1) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(7, 1)
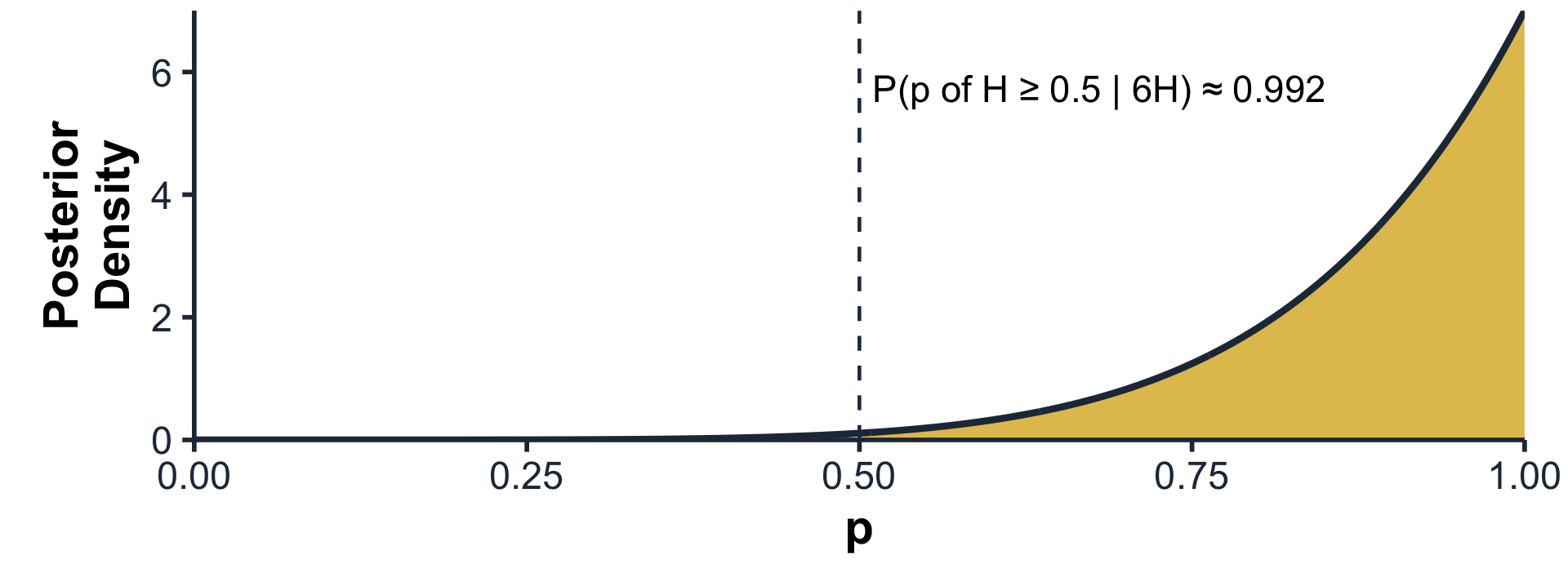
Bayesian Framework: P(Hypothesis | Data)

Prior: Beta(1, 1) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(7, 1)
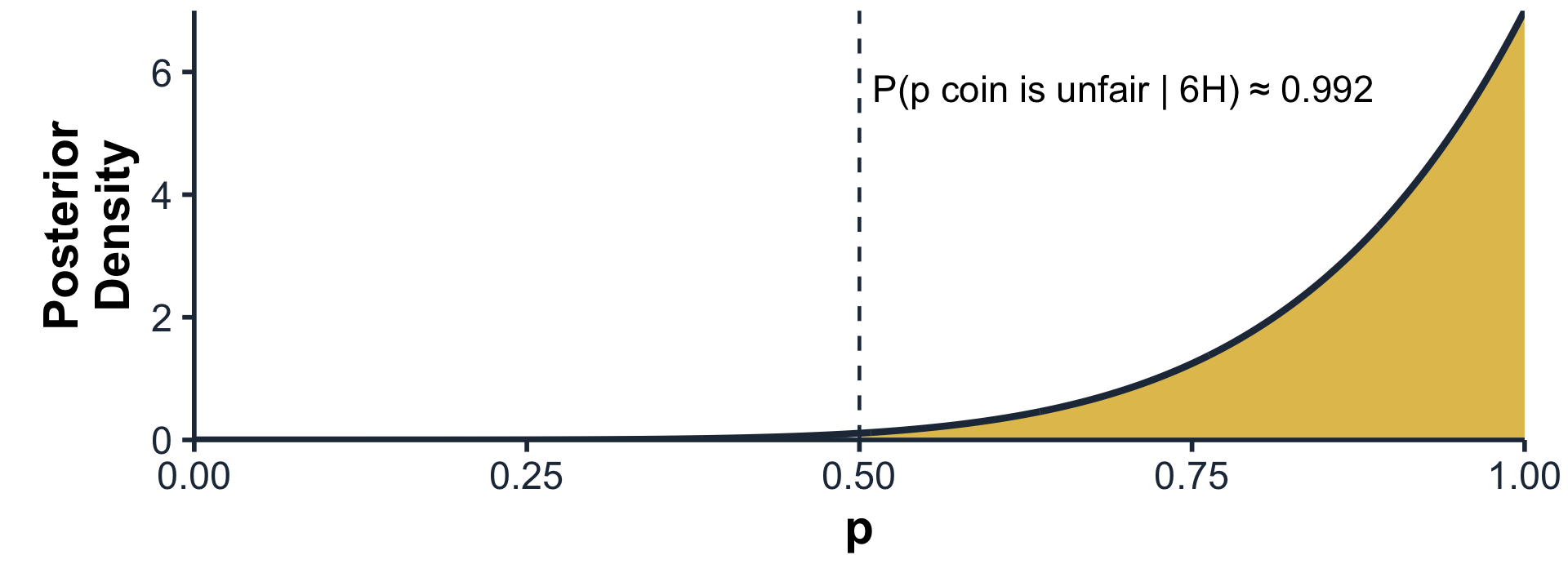
Bayesian Framework: P(Hypothesis | Data)

Prior: Beta(1, 1) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(7, 1)
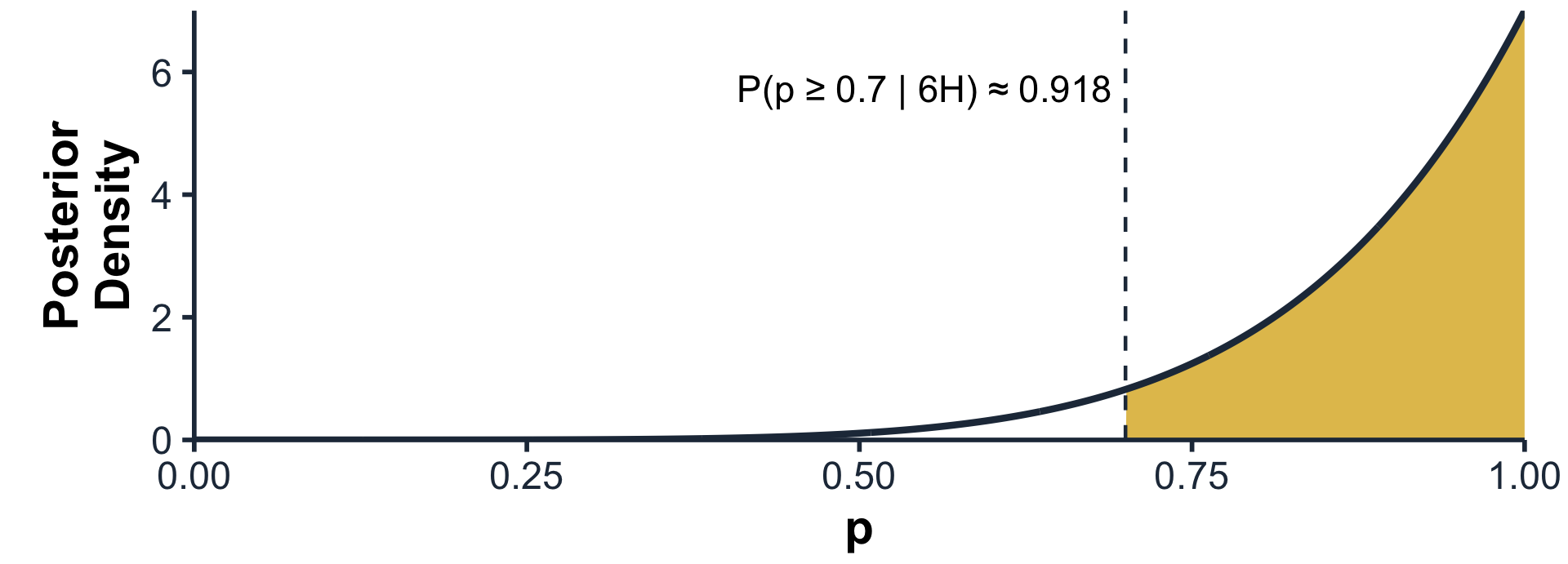
Bayesian Framework: P(Hypothesis | Data)

Prior: Beta(1, 1) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(7, 1)
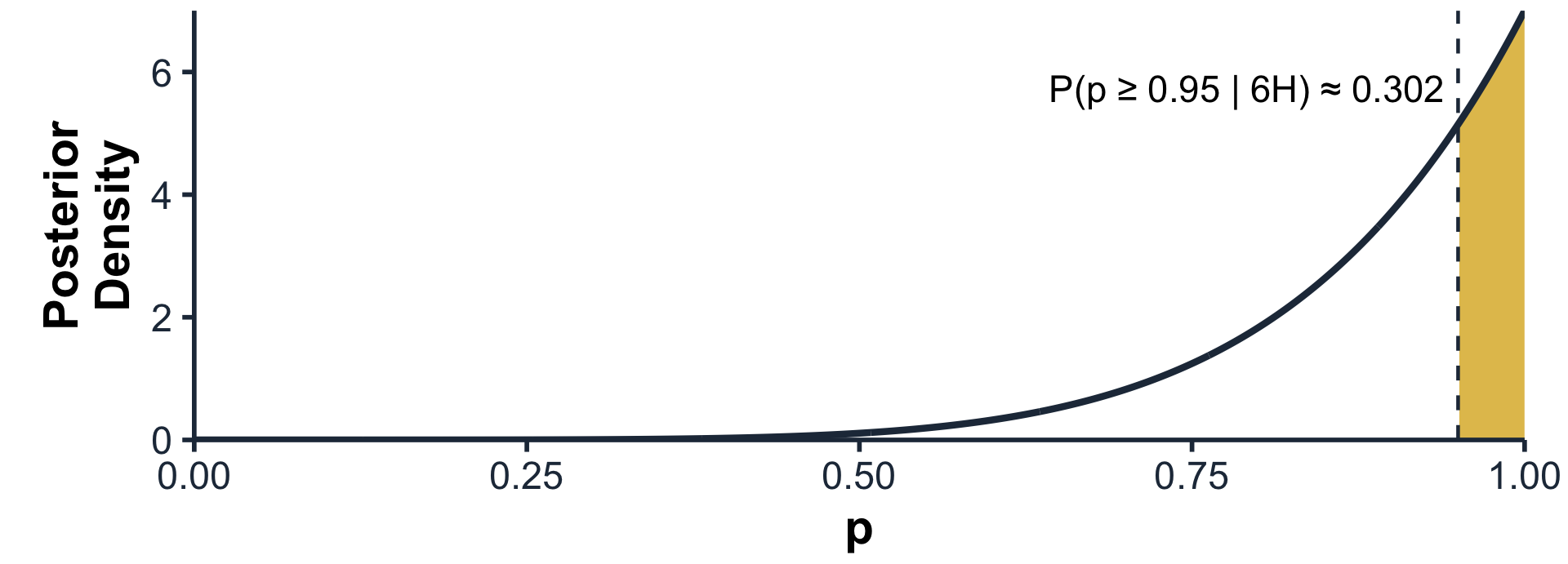
Bayesian Framework: The Importance of The Prior

Prior: Beta(21, 21) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(27, 21)
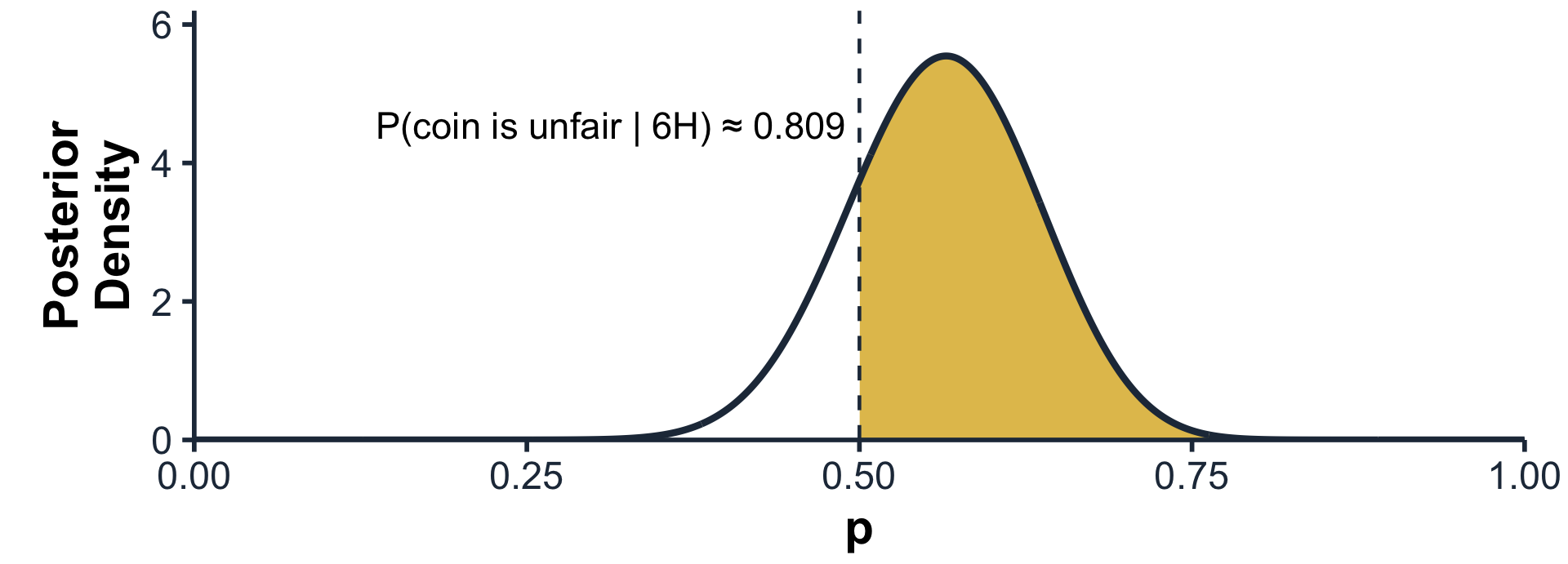
Bayesian Framework: The Effect of A “Strong” Prior

Prior: Beta(1001, 1001) Likelihood: P(6 heads | p) = p⁶
Posterior ∝ Prior × Likelihood:
P(p | 6H) ∝ P(p) · P(6H | p) ⇒ Posterior: Beta(1007, 1001)
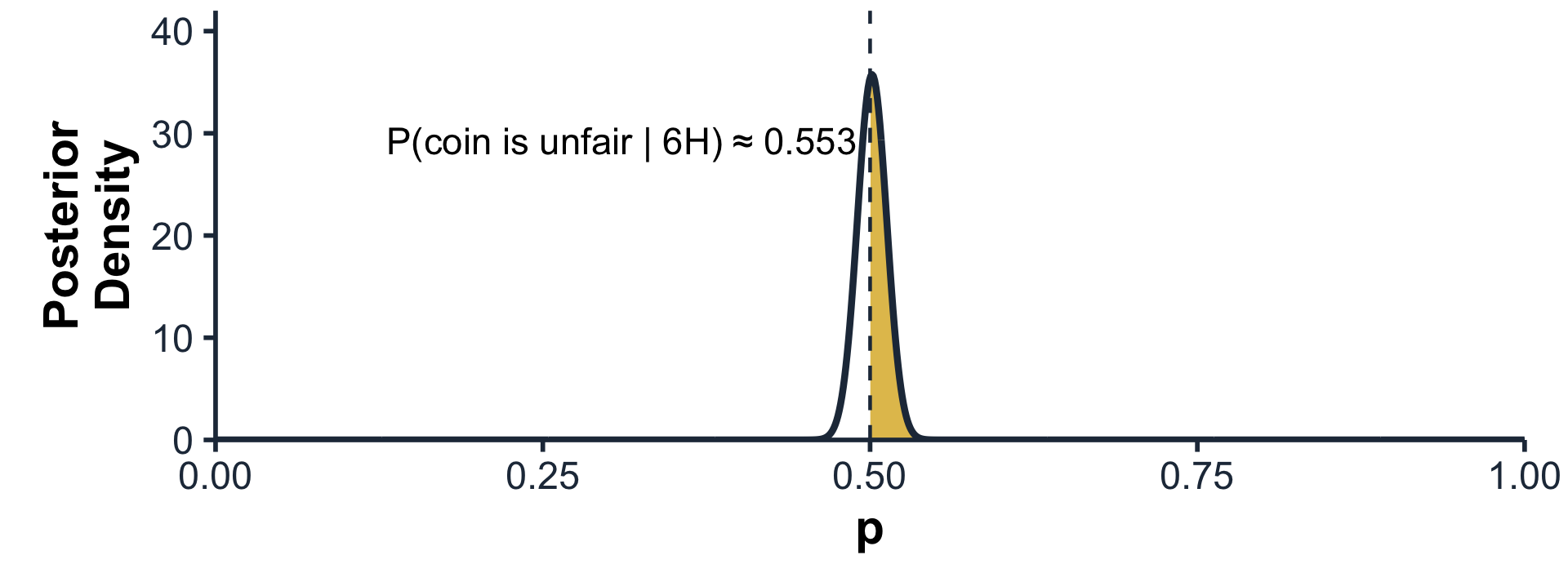
Two Frameworks For Interpreting Evidence
Frequentist Framework
Probability describes long-run behavior of data
Focus:
• How unusual are the data if no effect exists?
Typical outputs:
• p-values
• confidence intervals
Interpretation requires care:
A p-value does not tell us the probability a hypothesis is true
Bayesian Framework
Probability describes plausibility of hypotheses
Core idea:
Prior beliefs are updated by data
Key output:
• Posterior distribution
Range of plausible values for the effect
Interpretation is direct:
Probability statements apply to the quantity of interest
Two Frameworks For Interpreting Evidence
Key distinction:
• Frequentism evaluates how surprising the data are
• Bayesian analysis estimates how plausible different effect sizes are
KEYNOTE-630
Frequentist Analysis of KEYNOTE-630
Kaplan-Meier curves and Cox proportional hazards estimate
Frequentist Analysis of KEYNOTE-630
Kaplan-Meier curves and Cox proportional hazards estimate
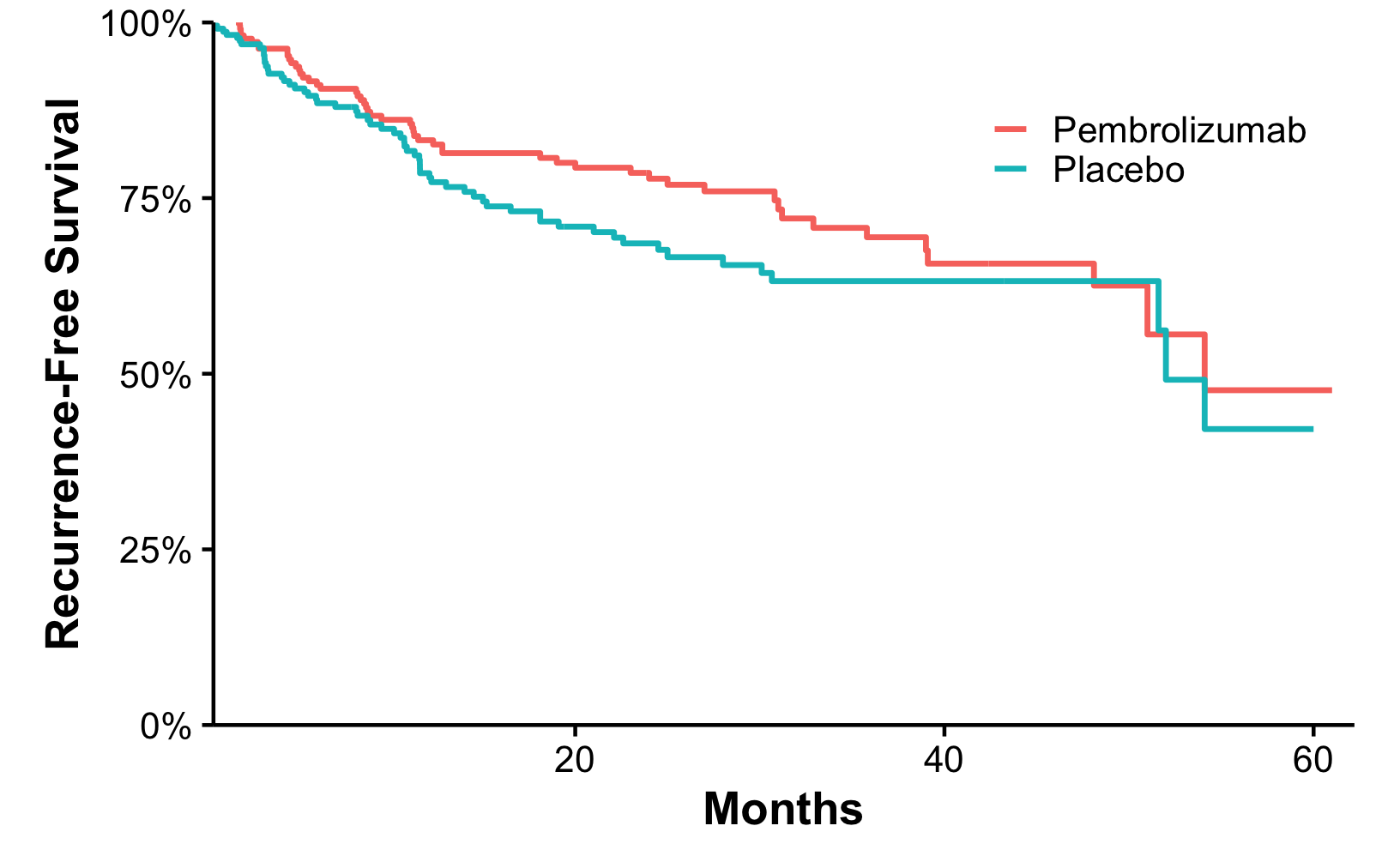
Frequentist Analysis of KEYNOTE-630
Kaplan-Meier curves and Cox proportional hazards estimate
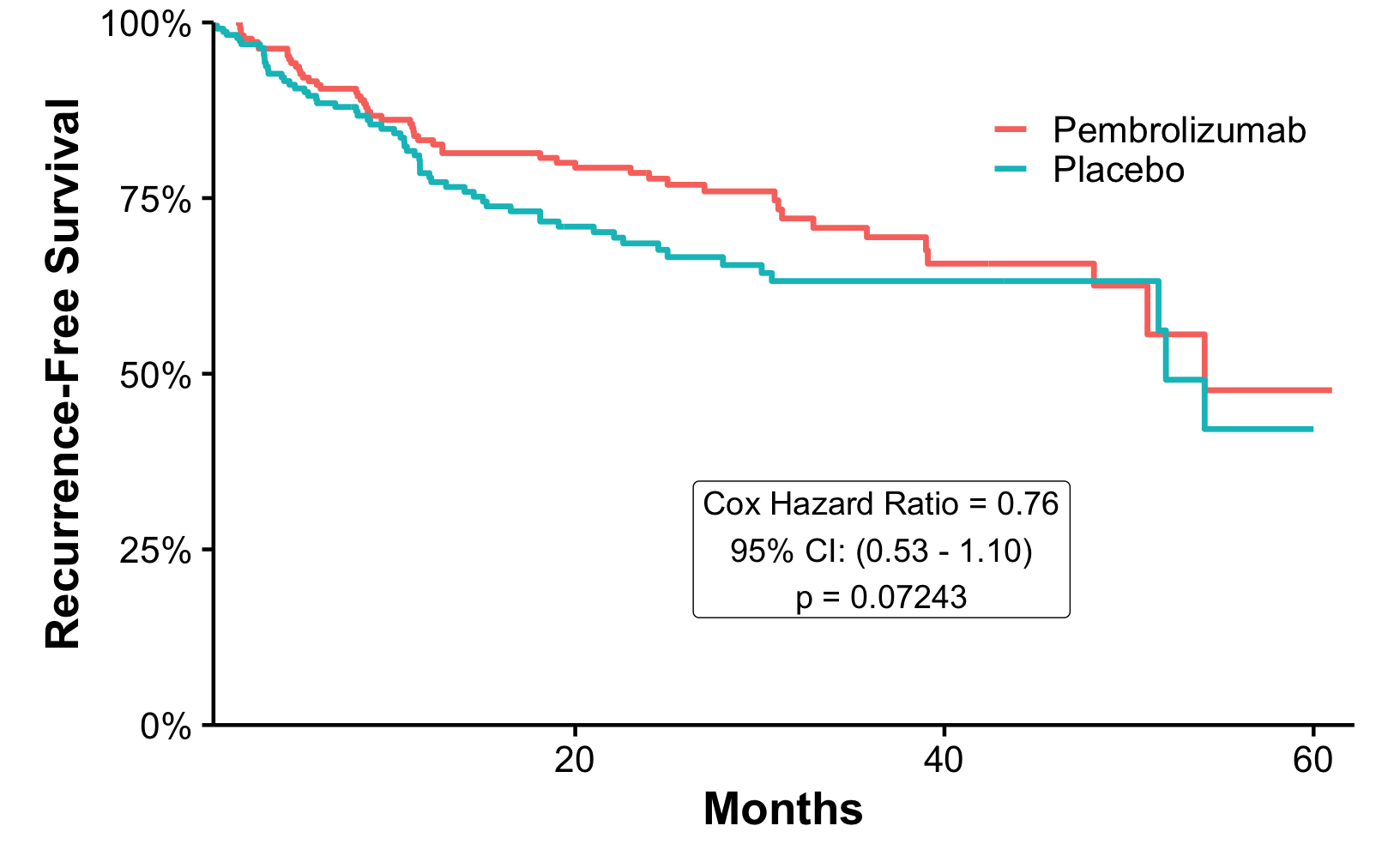
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
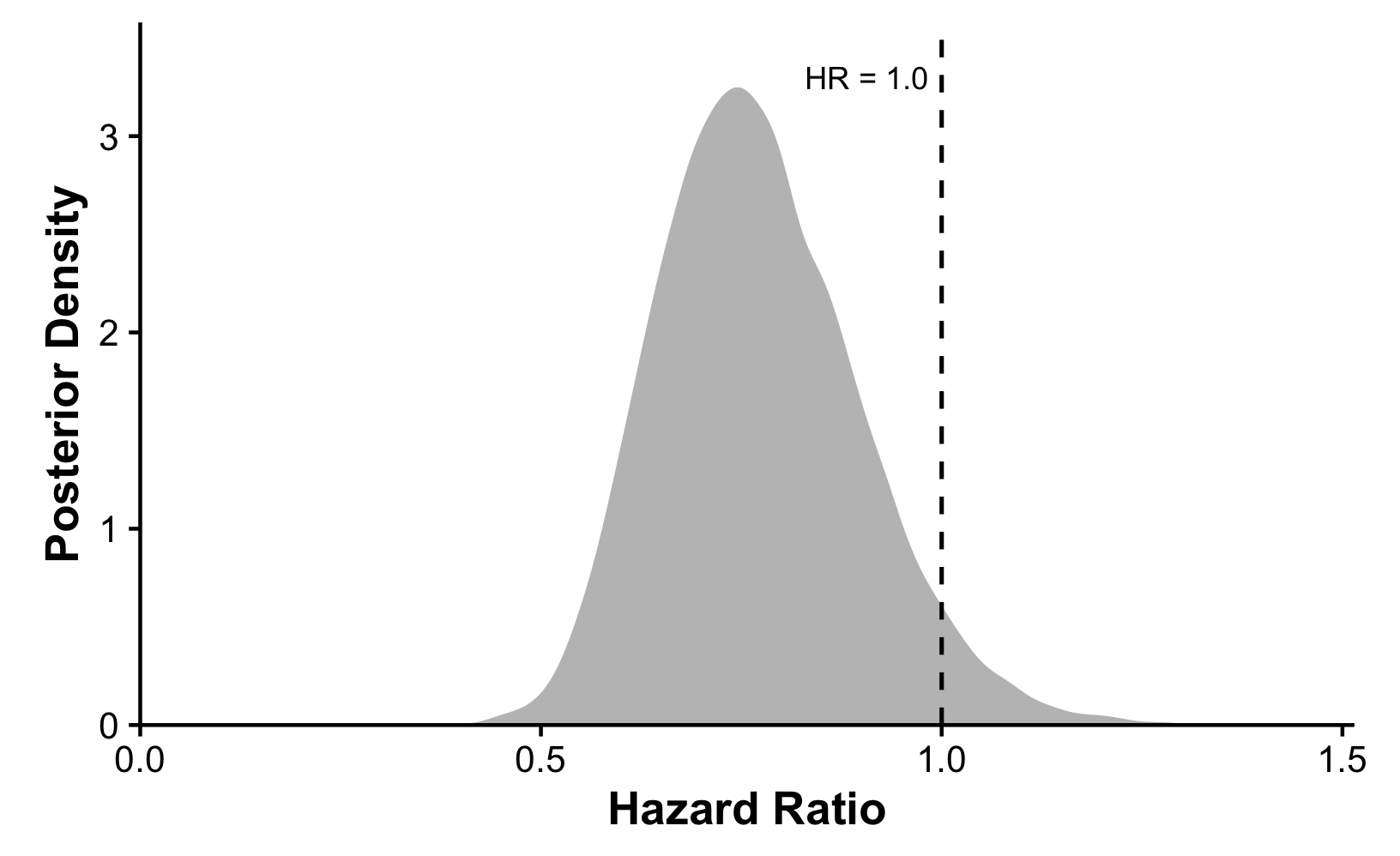
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
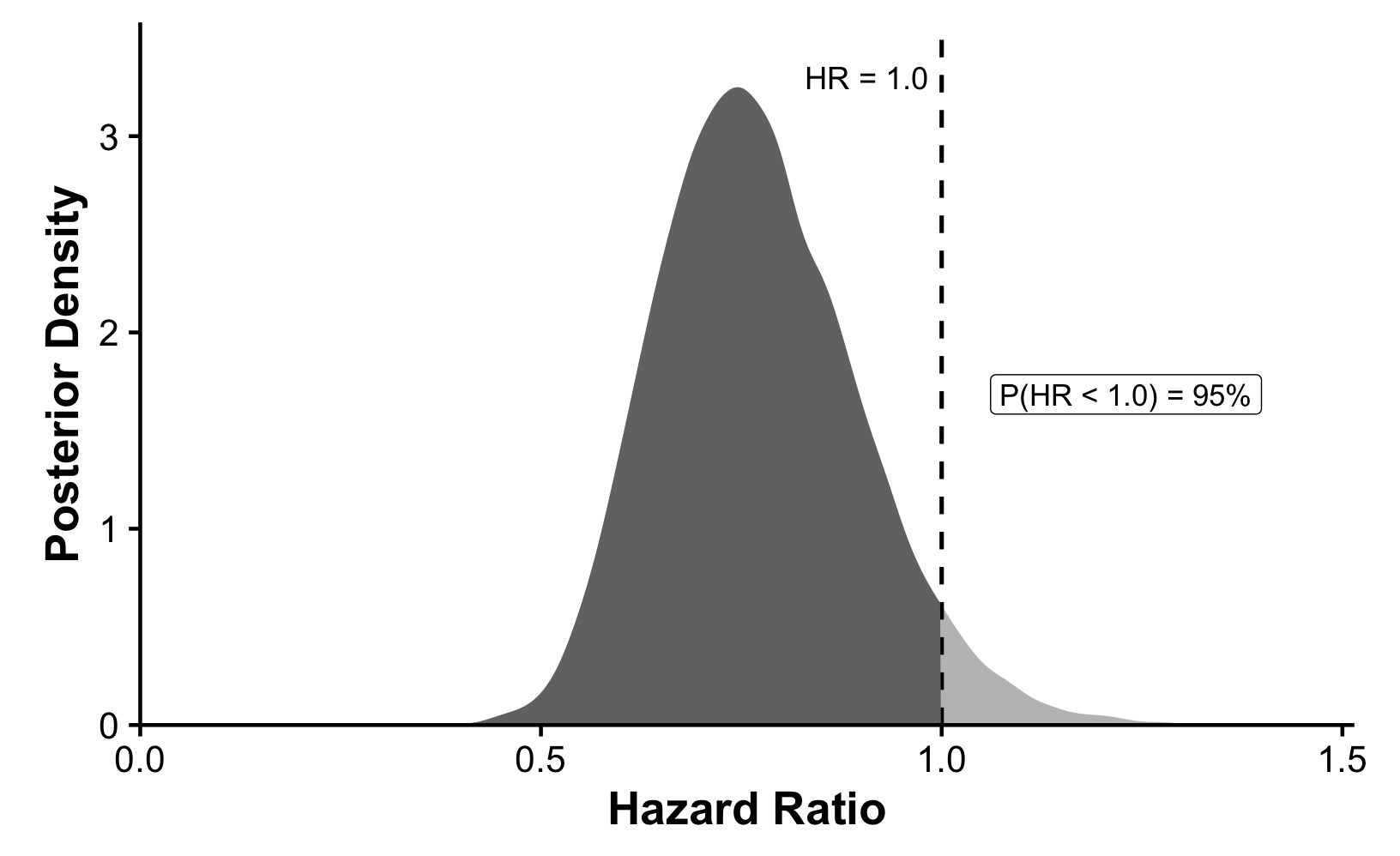
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
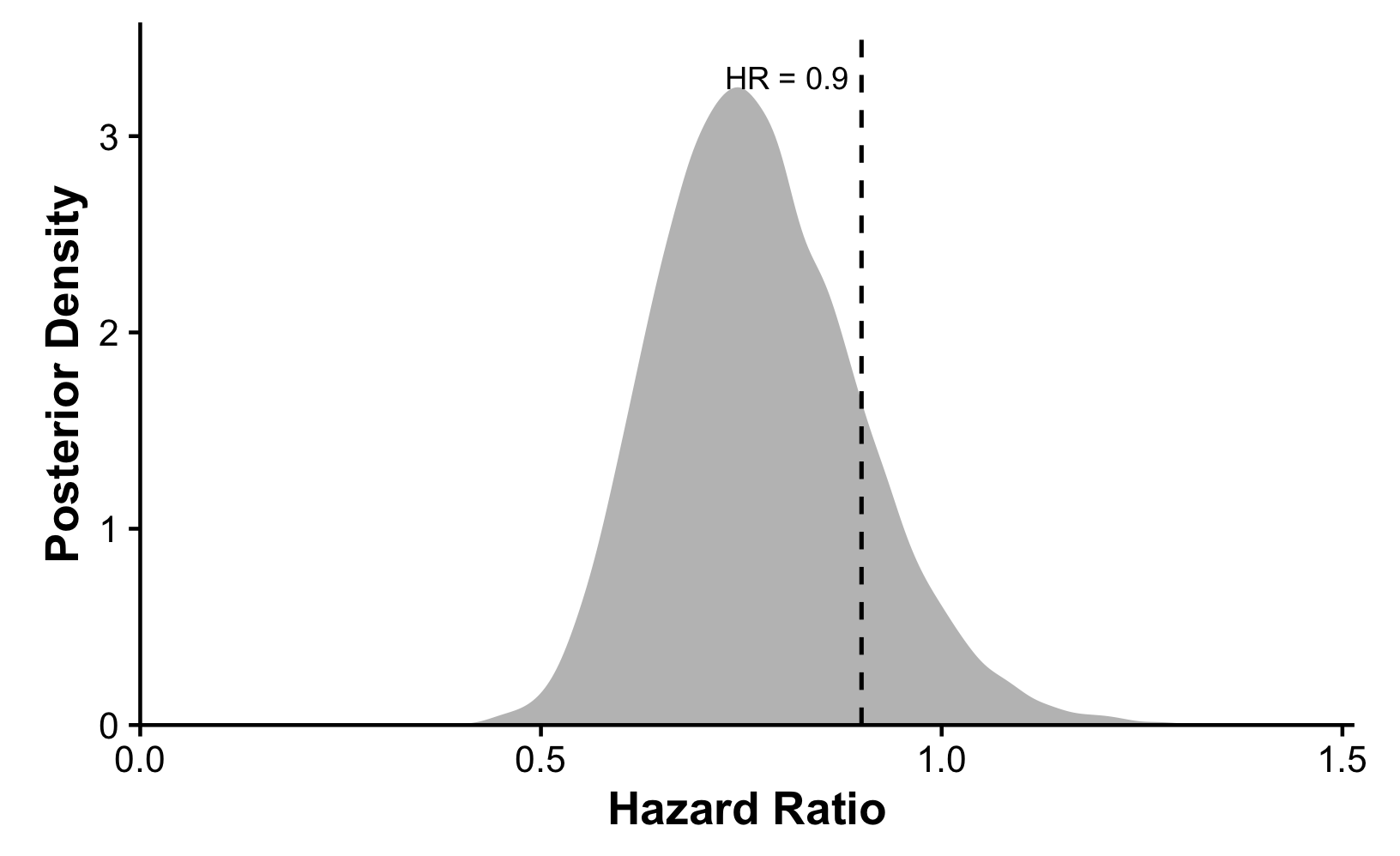
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
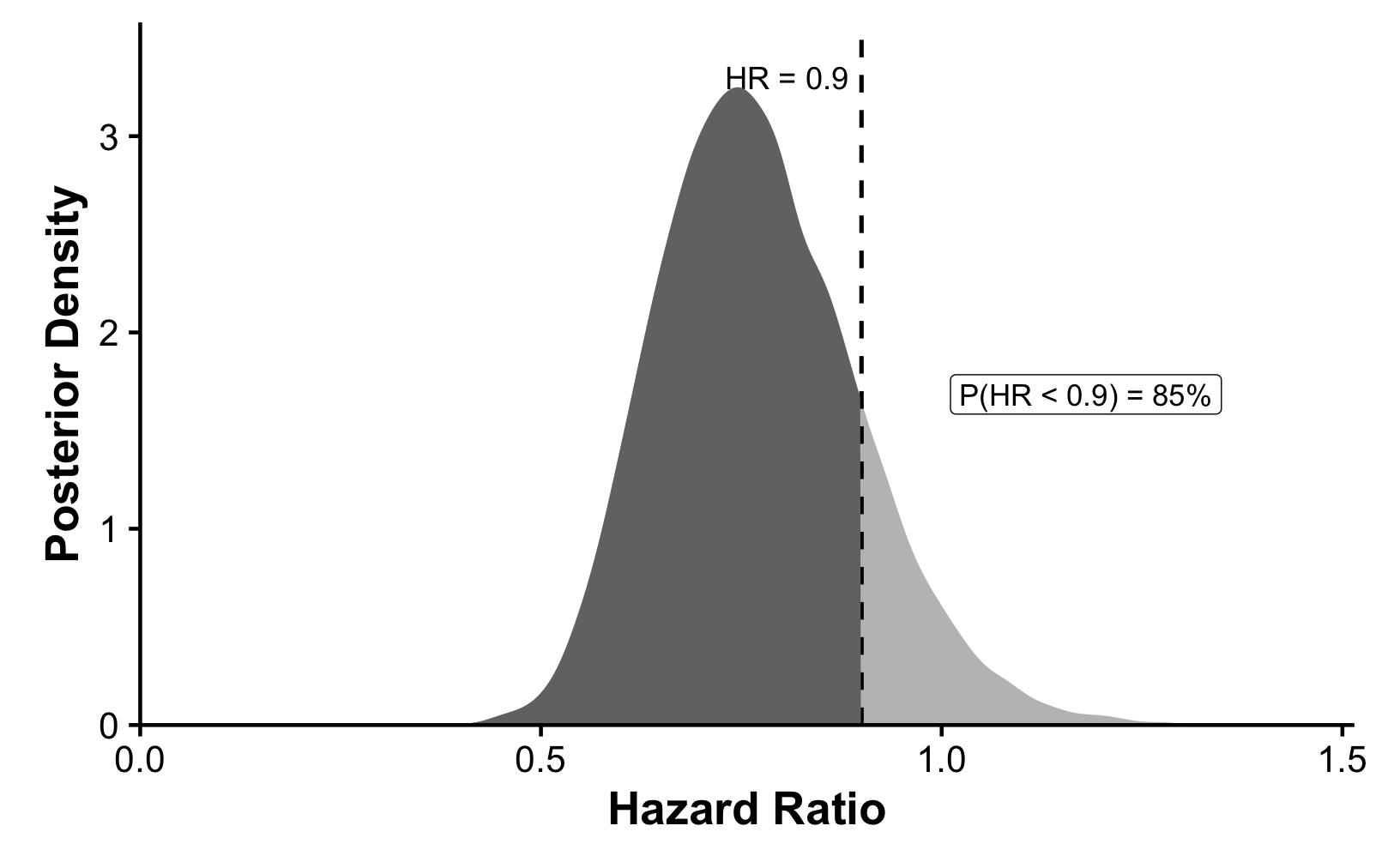
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
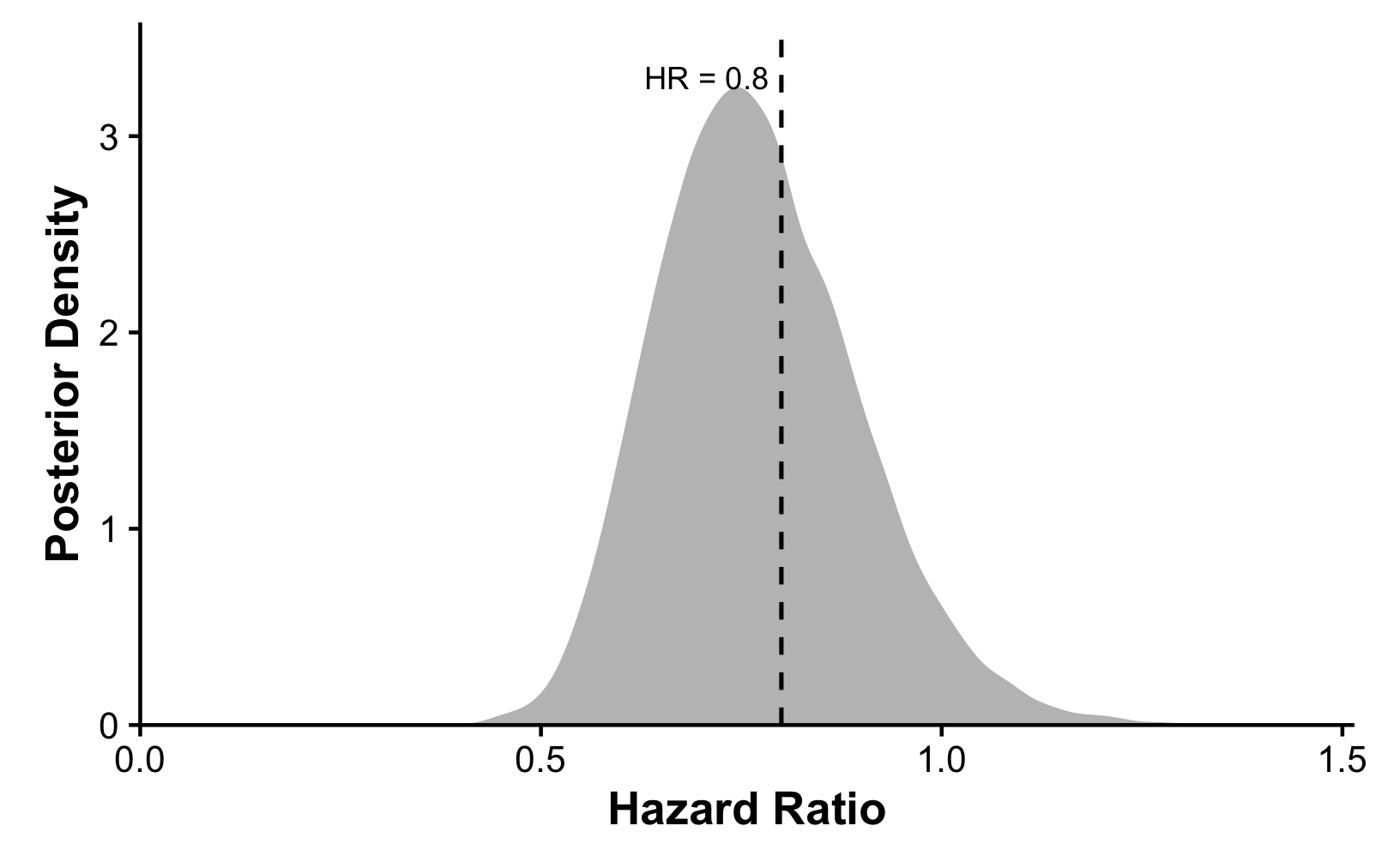
Bayesian Analysis of KEYNOTE-630
Posterior distribution of the hazard ratio
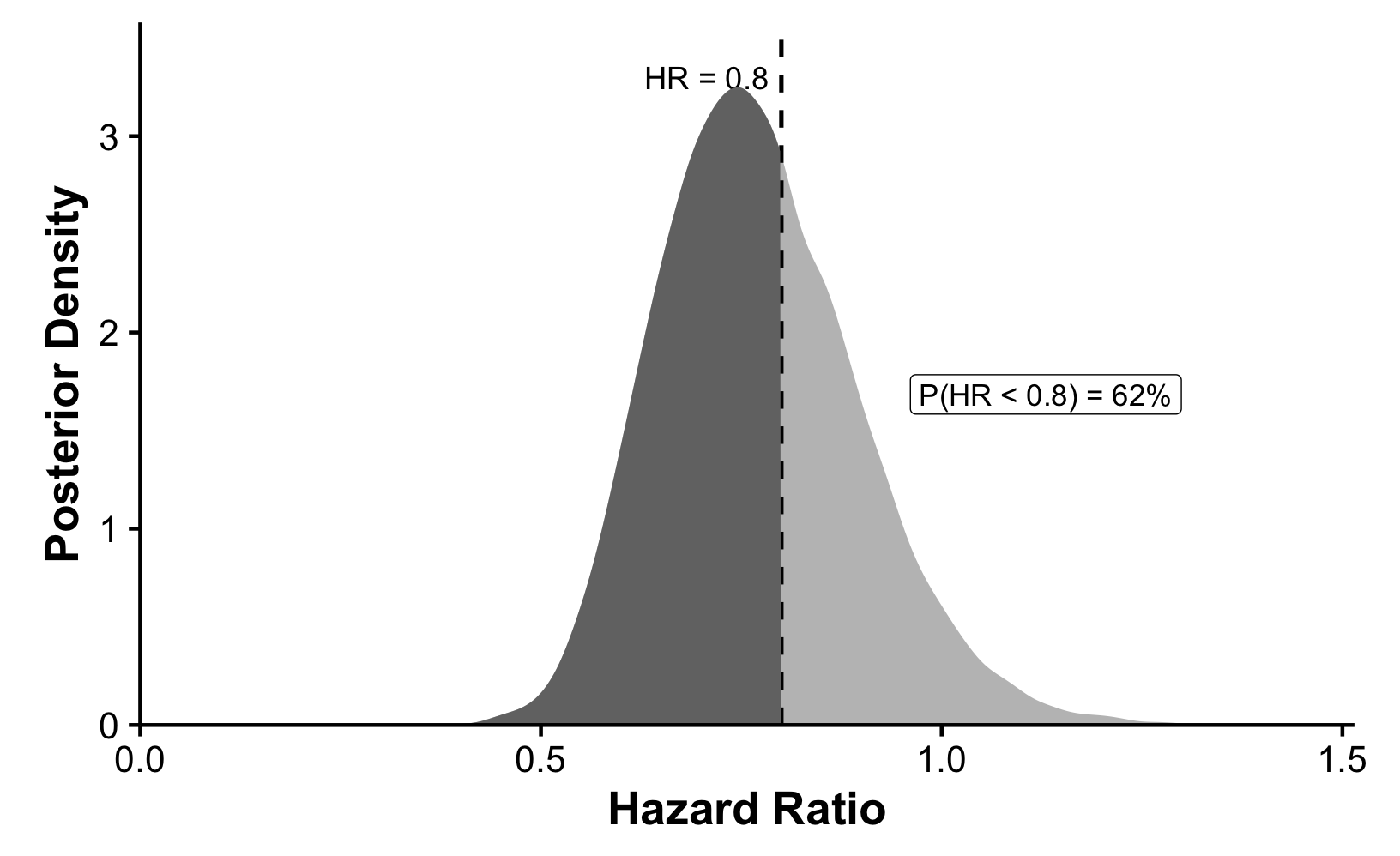
A Natural Question Follows
How Often Is Bayesian Analysis Actually Used in JAAD Papers?
A Natural Question Follows
How Often Is Bayesian Analysis Actually Used in JAAD Papers?
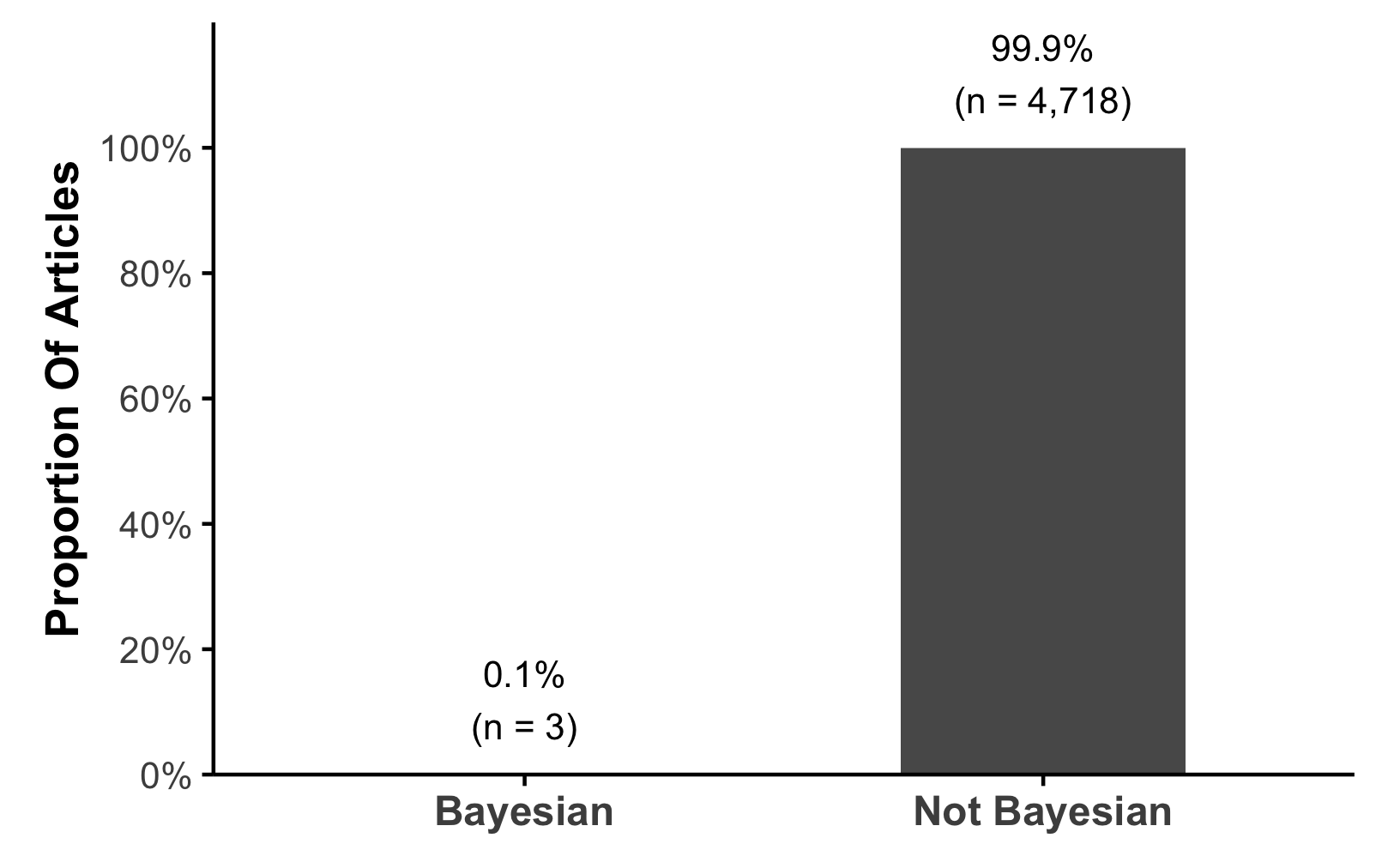
Summary
Core Ideas
• Researchers often have many reasonable ways to analyze the same data
• Results are often reduced to “statistically significant” or “not significant”
• This can hide how large an effect is and how uncertain we are
Insight
Statistical workflows influence
how results are interpreted
Key Takeaway
Interpretation should consider:
• Size of effect
• Uncertainty
• Clinical relevance
Not only whether p < 0.05
Observations From JAAD Review
• P-values are extremely common
• Adjustments for multiple testing are uncommon
• Preregistration is rare outside trials
• Bayesian approaches are rarely used
Study Limitations
Text-based extraction
- Some statistical information may not appear explicitly in text
- Figures and supplements may contain additional results
Operational definitions
- Identification of multiplicity and preregistration depends on detectable language
- Some analytic decisions may not be fully documented in publications
Corpus scope
- Focused on one high-impact dermatology journal
- Patterns may differ across journals
Analytic search space is not directly observable
- Reported tests represent only a portion of possible analytic pathways
Aligning Methods With Scientific Questions
Improving how we interpret evidence under analytic flexibility
Study Planning
• Clearly Define Hypotheses
• Distinguish Confirmation vs Exploration
• Embrace Exploratory Analysis
• Preregister When Feasible
Aligning Methods With Scientific Questions
Improving how we interpret evidence under analytic flexibility
Study Planning
• Clearly Define Hypotheses
• Distinguish Confirmation vs Exploration
• Embrace Exploratory Analysis
• Preregister When Feasible
Analysis Strategy
• Acknowledge Analytic Flexibility
• Consider implications of multiple comparisons
• Multiverse Analysis
Aligning Methods With Scientific Questions
Improving how we interpret evidence under analytic flexibility
Study Planning
• Clearly Define Hypotheses
• Distinguish Confirmation vs Exploration
• Embrace Exploratory Analysis
• Preregister When Feasible
Analysis Strategy
• Acknowledge Analytic Flexibility
• Consider implications of multiple comparisons
• Multiverse Analysis
Interpretation
• Avoid Rigid Significant / Non-Significant Framing
Bayesian Perspective
Bayesian Methods Can Provide:
• Direct Probability Statements
• Continuous evidence rather than binary thresholds
Core Principle
Statistical Methods Should Support Scientific Understanding Not Replace It With Threshold-Based Decisions
Thank You
Questions & Discussion

MGBCI